library(hmisindia)
library(dplyr)
library(ggplot2)
library(tidyr)
library(DT)
data_available <- tryCatch(
{
pq <- hmisindia::get_parquet_path()
nzchar(pq) && file.exists(pq)
},
error = function(e) FALSE
)
knitr::opts_chunk$set(eval = data_available)The hmisindia package provides access to India’s Health
Management Information System (HMIS) data: 840+ health indicators across
36 states/UTs from April 2008 to December 2024, with district-level data
for 700+ districts.
Browsing indicators
Find what’s available with list_indicators() and
search_indicators():
# All state-level indicators
list_indicators()
#> # A tibble: 840 × 4
#> canonical_name n_yearmon first_month last_month
#> <chr> <dbl> <chr> <chr>
#> 1 Above 10 years to below 19 years 24 Apr 2023 Mar 2025
#> 2 Above 5 years to below 10 years 24 Apr 2023 Mar 2025
#> 3 Adult above >19 years 24 Apr 2023 Mar 2025
#> 4 Albendazole 400 mg tablet 96 Apr 2017 Mar 2025
#> 5 Amoxycillin (Paediatrics Antibiotics) 24 Apr 2023 Mar 2025
#> 6 Antepartum (Macerated) Still Birth 24 Apr 2023 Mar 2025
#> 7 Average kilometres travelled by ALS during … 24 Apr 2023 Mar 2025
#> 8 Average kilometres travelled by BLS during … 24 Apr 2023 Mar 2025
#> 9 Average number of trips per day by ALS duri… 24 Apr 2023 Mar 2025
#> 10 Average number of trips per day by BLS duri… 24 Apr 2023 Mar 2025
#> # ℹ 830 more rows
# Search by keyword
search_indicators("malaria")
#> # A tibble: 14 × 4
#> canonical_name n_yearmon first_month last_month
#> <chr> <dbl> <chr> <chr>
#> 1 Childhood Diseases - Malaria 96 Apr 2017 Mar 2025
#> 2 Inpatient - Malaria 96 Apr 2017 Mar 2025
#> 3 Malaria (Microscopy Tests) - Mixed test pos… 24 Apr 2023 Mar 2025
#> 4 Malaria (Microscopy Tests) - Plasmodium Fal… 96 Apr 2017 Mar 2025
#> 5 Malaria (Microscopy Tests) - Plasmodium Viv… 96 Apr 2017 Mar 2025
#> 6 Malaria (RDT) - Plamodium Falciparum test p… 96 Apr 2017 Mar 2025
#> 7 Number of blood smears examined for Malaria 204 Apr 2008 Mar 2025
#> 8 Number of cases of Adolescent or Adult deat… 108 Apr 2008 Mar 2017
#> 9 Number of cases of Malaria reported in chil… 108 Apr 2008 Mar 2017
#> 10 Number of Deaths due to Malaria - Plasmodiu… 96 Apr 2017 Mar 2025
#> 11 Number of Deaths due to Malaria - Plasmodiu… 96 Apr 2017 Mar 2025
#> 12 Out of blood smears examined for malaria, n… 108 Apr 2008 Mar 2017
#> 13 Out of blood smears examined for malaria, n… 108 Apr 2008 Mar 2017
#> 14 RDT conducted for Malaria 96 Apr 2017 Mar 2025
# District-level indicators
search_indicators("live birth", geography = "district")
#> # A tibble: 4 × 4
#> canonical_name n_yearmon first_month last_month
#> <chr> <int> <chr> <chr>
#> 1 Number of female live births 204 Apr 2008 Mar 2025
#> 2 Number of live births among Syphilis Seropos… 24 Apr 2023 Mar 2025
#> 3 Number of male live births 204 Apr 2008 Mar 2025
#> 4 Total number of male and female live births … 108 Apr 2008 Mar 2017Querying data
get_hmis() queries indicator data by substring match
(default), regex, or fuzzy match:
# All data for an indicator
births <- get_hmis("Number of female live births",
category = "Total",
sector = "Total"
)
datatable(births, options = list(pageLength = 10, scrollX = TRUE))Filter by state, time period, and more:
Time series plots
plot_time_series() creates line plots colored by state
(or district, indicator, etc.):
plot_time_series(births_subset, y_label = "Number of live births (female)")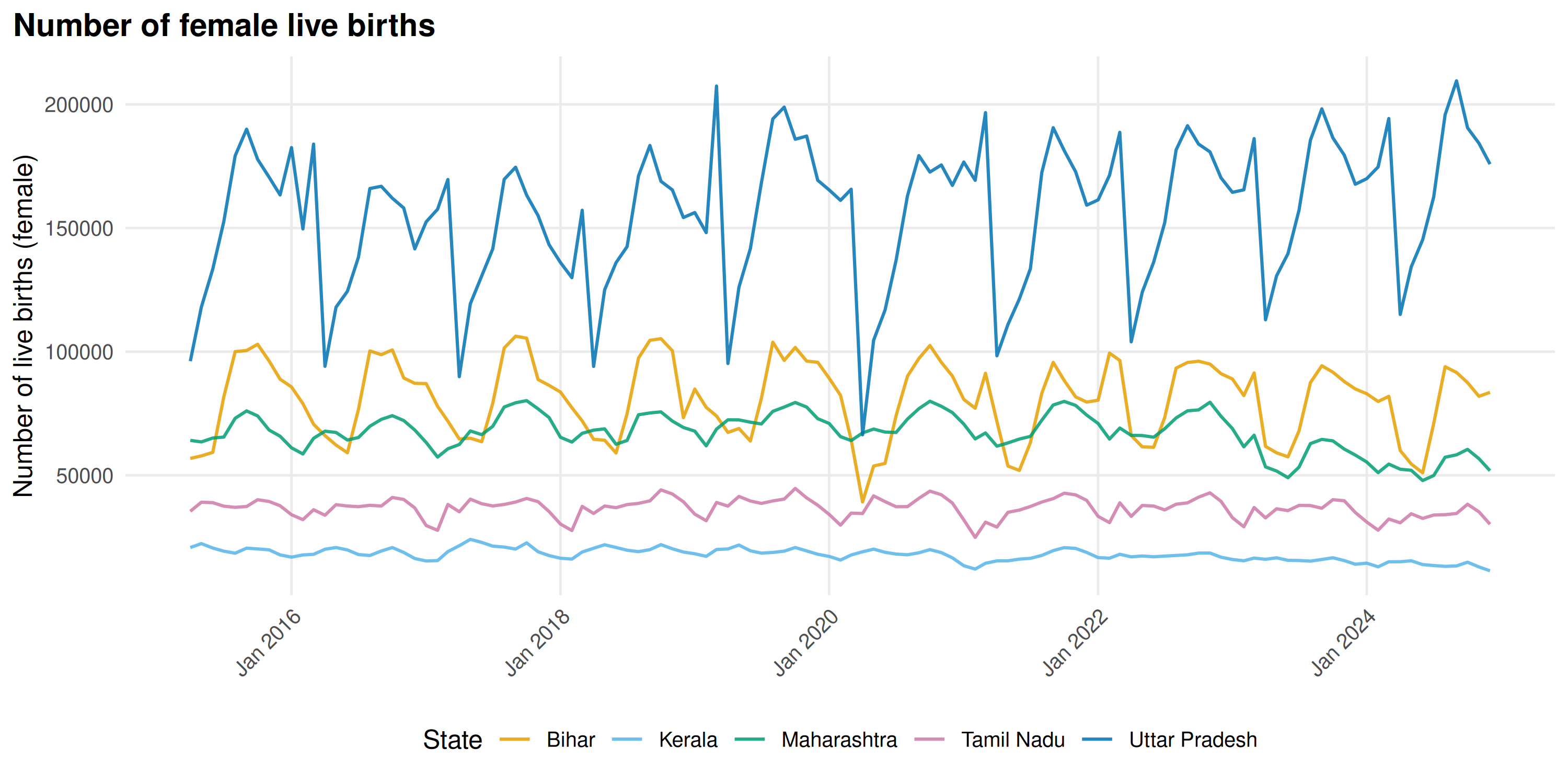
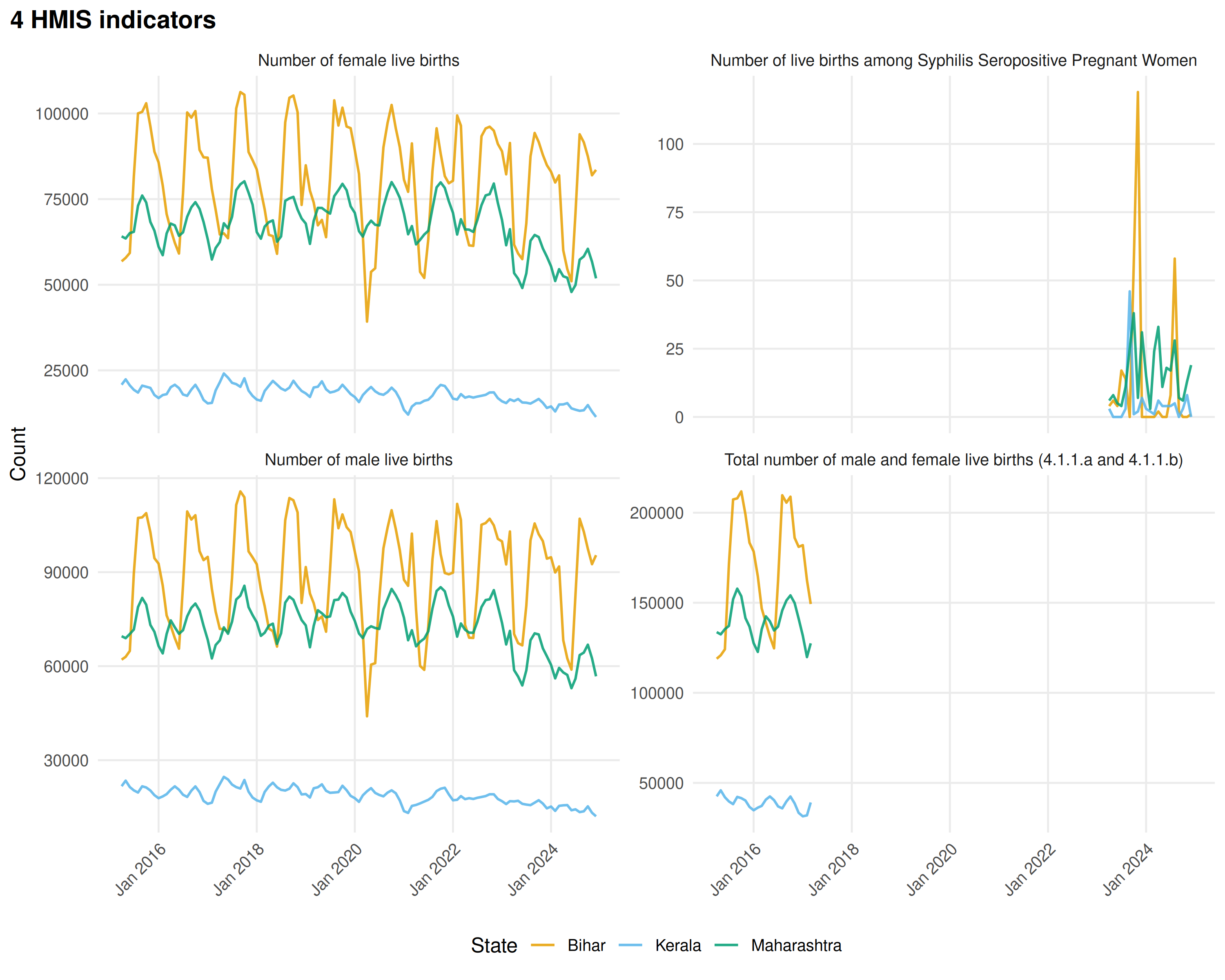
Compare multiple indicators by faceting:
birth_indicators <- get_hmis("live births",
state = c("Bihar", "Kerala", "Maharashtra"),
category = "Total",
sector = "Total",
from = "Apr 2015",
to = "Dec 2024"
)
plot_time_series(birth_indicators,
facet = "canonical_name",
y_label = "Count"
)
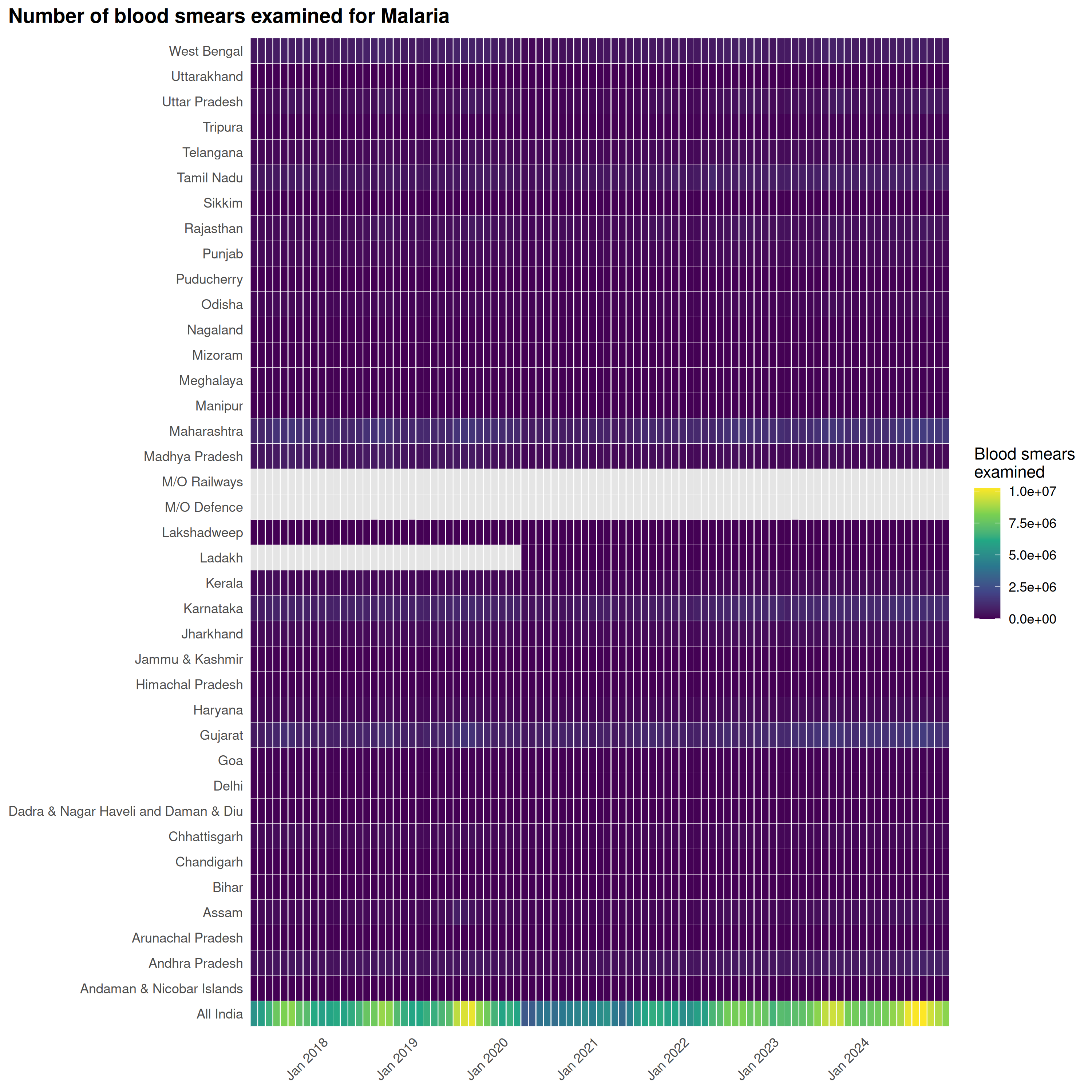
Heatmaps
plot_heatmap() shows values across states and time:
malaria <- get_hmis("Number of blood smears examined for Malaria",
category = "Total",
sector = "Total",
from = "Apr 2017",
to = "Dec 2024"
)
plot_heatmap(malaria,
legend_title = "Blood smears\nexamined",
palette = "viridis"
)
Scatter plots
plot_scatter() compares two variables. Here we compare
male vs female live births:
male_births <- get_hmis("Number of male live births",
category = "Total",
sector = "Total",
from = "Apr 2015",
to = "Dec 2024"
)
female_births <- get_hmis("Number of female live births",
category = "Total",
sector = "Total",
from = "Apr 2015",
to = "Dec 2024"
)
# Merge the two indicators into wide format
scatter_data <- inner_join(
male_births |> select(state, monyear, male = value),
female_births |> select(state, monyear, female = value),
by = c("state", "monyear")
)
plot_scatter(scatter_data,
x = "male",
y = "female",
x_label = "Male live births",
y_label = "Female live births",
title = "Male vs female live births across states"
)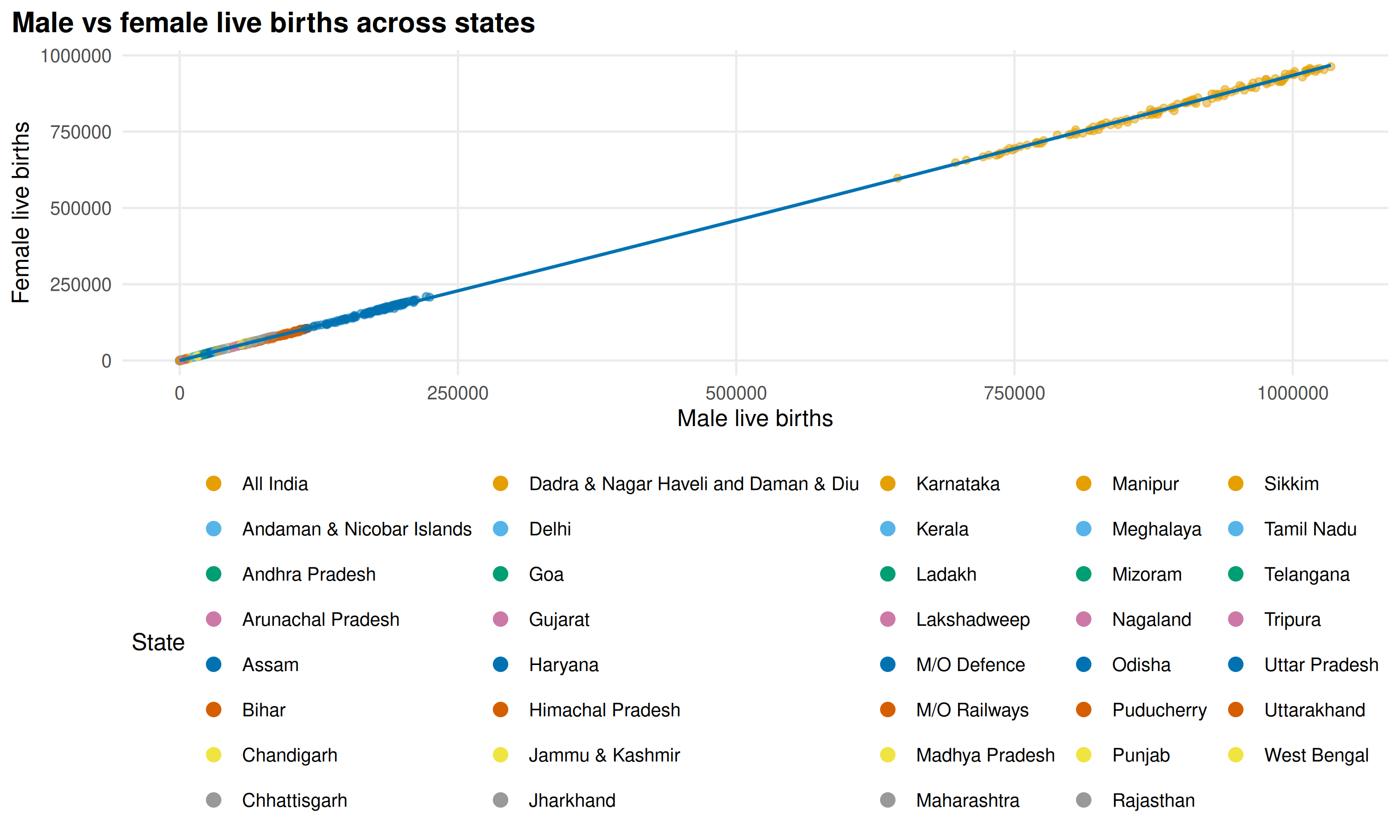
Sex ratios at birth are consistent across states and time.
Periodicity detection
detect_periodicity() tests for seasonal patterns using
spectral analysis and autocorrelation:
kerala_births <- get_hmis("Number of female live births",
state = "Kerala",
category = "Total",
sector = "Total"
)
prd <- detect_periodicity(kerala_births)
print(prd)
#> Periodicity analysis (n = 204 observations)
#> Detrended: TRUE
#> Dominant period: 6.2 observations
#> Spectral power concentration: 6.4 %
#> ACF at dominant lag: 0.305 (threshold: 0.137 )
#>
#> Top frequencies:
#> period power
#> 6.17 19215478
#> 6.00 19009942
#> 5.84 18639115
#> 6.35 18599844
#> 5.68 18036973
plot(prd)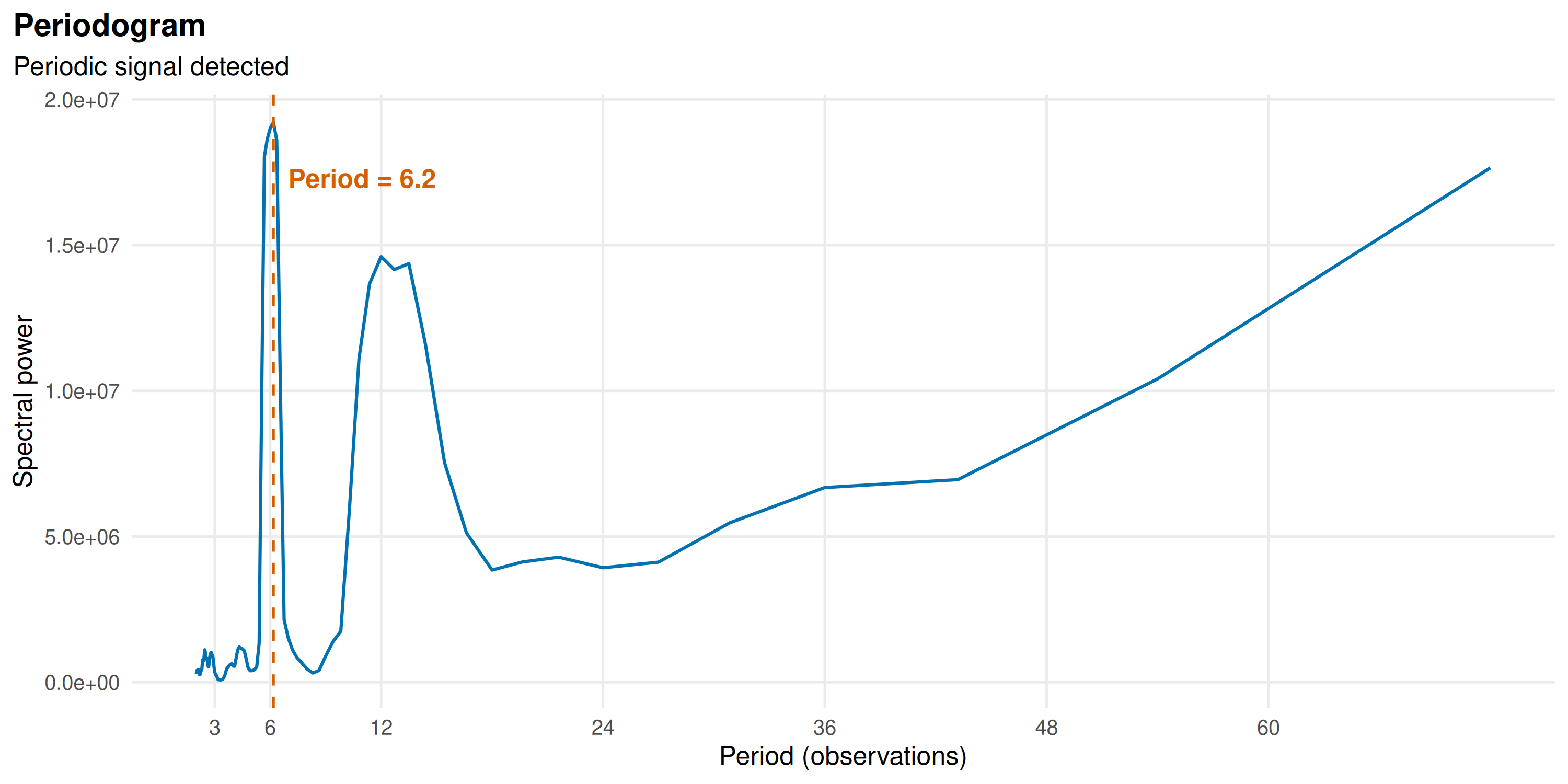
A spectral peak at 12 months indicates an annual cycle.
Decomposing seasonal components
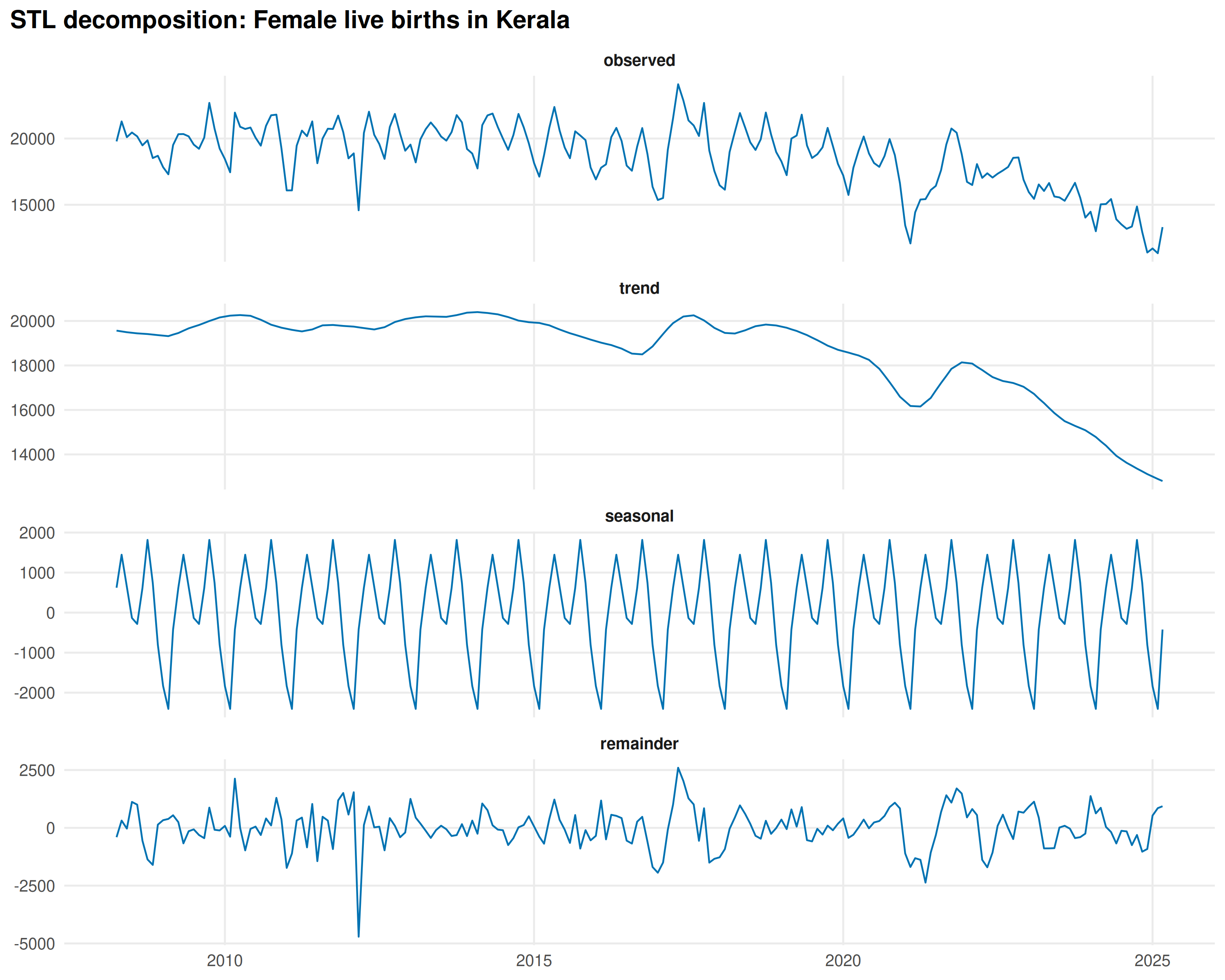
decompose_series() separates trend, seasonal, and
residual components using STL decomposition:
decomp <- decompose_series(kerala_births)
decomp_long <- decomp |>
pivot_longer(
cols = c(observed, trend, seasonal, remainder),
names_to = "component",
values_to = "val"
) |>
mutate(component = factor(component,
levels = c("observed", "trend", "seasonal", "remainder")
))
ggplot(decomp_long, aes(date, val)) +
geom_line(color = "#0072B2", linewidth = 0.5) +
facet_wrap(~component, ncol = 1, scales = "free_y") +
labs(
x = NULL, y = NULL,
title = "STL decomposition: Female live births in Kerala"
) +
theme_minimal(base_size = 12) +
theme(
strip.text = element_text(face = "bold"),
panel.grid.minor = element_blank(),
plot.title = element_text(face = "bold"),
plot.title.position = "plot"
)
District-level data
Query district data by setting
geography = "district":
# List districts in a state
list_districts(state = "Bihar")
#> # A tibble: 38 × 2
#> state district
#> <chr> <chr>
#> 1 Bihar Araria
#> 2 Bihar Arwal
#> 3 Bihar Aurangabad
#> 4 Bihar Banka
#> 5 Bihar Begusarai
#> 6 Bihar Bhagalpur
#> 7 Bihar Bhojpur
#> 8 Bihar Buxar
#> 9 Bihar Darbhanga
#> 10 Bihar Gaya
#> # ℹ 28 more rows
# District-level malaria data
bihar_malaria <- get_hmis("Number of blood smears examined for Malaria",
geography = "district",
state = "Bihar",
category = "Total",
sector = "Total",
from = "Apr 2020",
to = "Dec 2024"
)
# Show top 8 districts by volume
top_districts <- bihar_malaria |>
group_by(district) |>
summarise(total = sum(value, na.rm = TRUE)) |>
slice_max(total, n = 8) |>
pull(district)
bihar_malaria |>
filter(district %in% top_districts) |>
plot_time_series(
color = "district",
y_label = "Blood smears examined",
title = "Malaria testing in Bihar's top 8 districts"
)District heatmaps
bihar_malaria |>
filter(district %in% top_districts) |>
plot_heatmap(
y_var = "district",
legend_title = "Blood smears\nexamined",
palette = "viridis",
title = "Malaria testing across Bihar districts"
)Mapping with geometry
Attach LGD boundary geometries for choropleth mapping. Requires the
sf package:
# Get annual average institutional deliveries by state
deliveries <- get_hmis("Number of Institutional Deliveries conducted",
category = "Total",
sector = "Total",
from = "Apr 2022",
to = "Dec 2024",
geometry = TRUE
)
# Aggregate to annual
deliveries_annual <- deliveries |>
sf::st_drop_geometry() |>
group_by(state) |>
summarise(total_deliveries = sum(value, na.rm = TRUE), .groups = "drop")
# Reattach geometry
deliveries_sf <- attach_geometry(deliveries_annual)
ggplot(deliveries_sf) +
geom_sf(aes(fill = total_deliveries / 1000), color = "white", linewidth = 0.2) +
scale_fill_viridis_c(option = "viridis", name = "Deliveries\n(thousands)") +
labs(title = "Institutional deliveries by state (Apr 2022 - Dec 2024)") +
theme_void(base_size = 12) +
theme(
plot.title = element_text(face = "bold"),
legend.position = "right"
)Summary
| Function | Purpose |
|---|---|
get_hmis() |
Query indicator data (state or district level) |
list_indicators() |
Browse available indicators |
search_indicators() |
Search indicators by keyword |
list_states() |
List all 36 states/UTs |
list_districts() |
List districts (with optional state filter) |
plot_time_series() |
Line plots of indicator values over time |
plot_heatmap() |
Heatmap of values across states/districts and time |
plot_scatter() |
Scatter plot comparing two variables |
detect_periodicity() |
Spectral analysis for seasonal patterns |
decompose_series() |
STL decomposition into trend + seasonal + remainder |
attach_geometry() |
Join LGD boundaries for mapping |
get_boundaries() |
Download state or district boundary geometries |