Identifying periodic indicators in HMIS data
Source:vignettes/periodic-indicators.Rmd
periodic-indicators.Rmd
library(hmisindia)
library(dplyr)
library(ggplot2)
library(tidyr)
library(DT)
data_available <- tryCatch(
{
pq <- hmisindia::get_parquet_path()
nzchar(pq) && file.exists(pq)
},
error = function(e) FALSE
)
knitr::opts_chunk$set(eval = data_available)Some HMIS indicators are seasonal: births peak in certain months, malaria tracks the monsoon. This scan covers all state-level indicators for periodicity at the national level.
Scanning all indicators
Each indicator with at least 36 months of coverage is aggregated to a
national monthly total, then tested with
detect_periodicity().
indicators <- list_indicators(min_months = 36)
pq_path <- get_parquet_path()
all_data <- arrow::open_dataset(pq_path) |>
dplyr::select(canonical_name, monyear, value) |>
collect()
national_all <- all_data |>
group_by(canonical_name, monyear) |>
summarise(value = sum(value, na.rm = TRUE), .groups = "drop")
# Filter to indicators with at least 36 months
ind_counts <- national_all |>
group_by(canonical_name) |>
summarise(n_months = n_distinct(monyear), .groups = "drop") |>
filter(n_months >= 36)
national_all <- national_all |>
semi_join(ind_counts, by = "canonical_name")
ind_names <- unique(national_all$canonical_name)
results <- lapply(ind_names, function(ind_name) {
tryCatch(
{
national <- national_all |> filter(canonical_name == ind_name)
if (nrow(national) < 36) {
return(NULL)
}
prd <- detect_periodicity(national)
data.frame(
canonical_name = ind_name,
n_months = prd$n,
is_periodic = prd$is_periodic,
dominant_period = round(prd$dominant_period, 1),
spectral_power_pct = round(prd$spectral_power * 100, 1),
acf_peak = round(prd$acf_peak, 3),
acf_threshold = round(prd$acf_threshold, 3),
stringsAsFactors = FALSE
)
},
error = function(e) NULL
)
})
scan_results <- do.call(rbind, Filter(Negate(is.null), results))
scan_results <- as_tibble(scan_results)Distribution of dominant periods
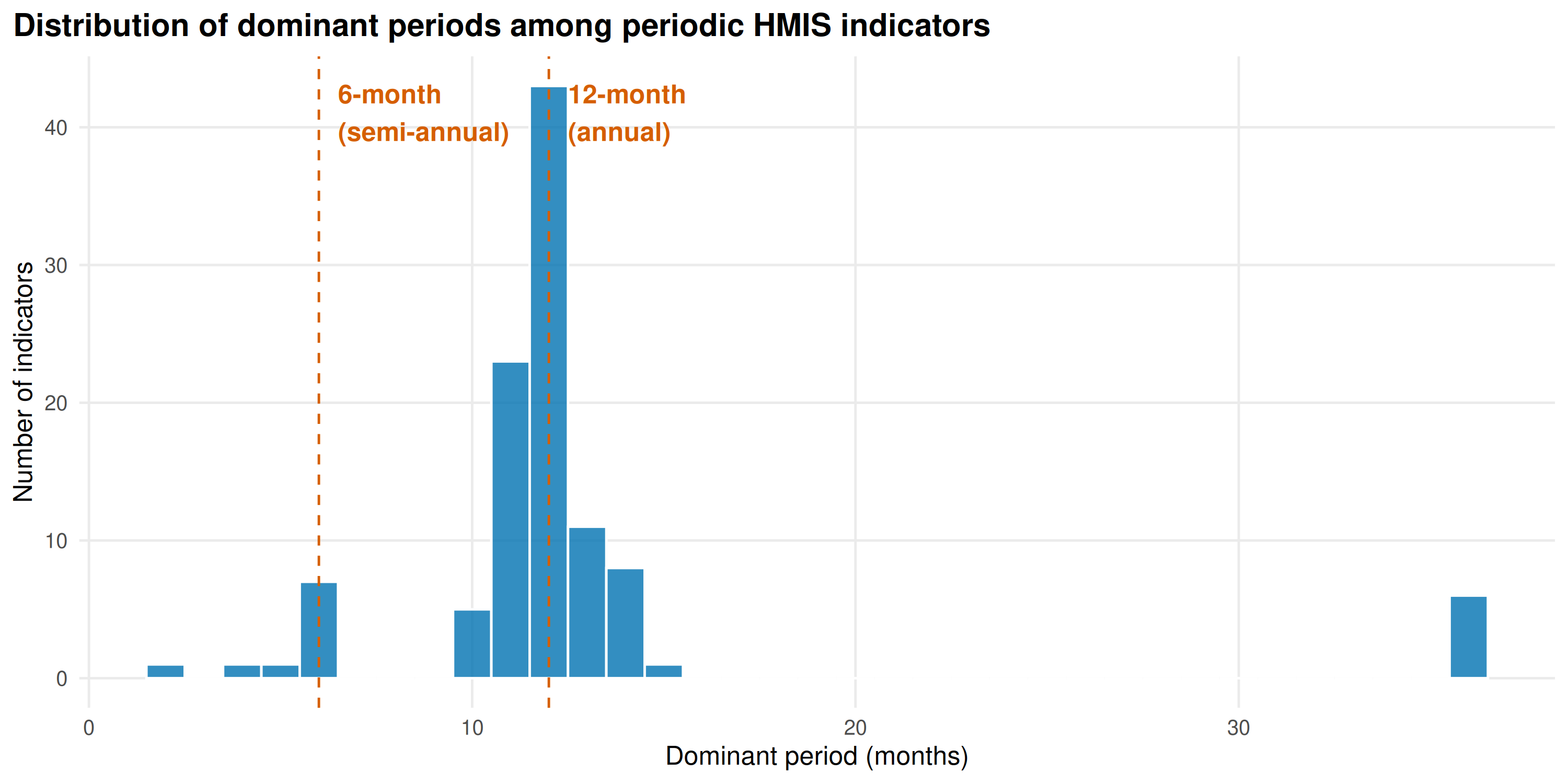
Cycle length distribution among periodic indicators:
periodic <- scan_results |> filter(is_periodic)
ggplot(periodic, aes(dominant_period)) +
geom_histogram(binwidth = 1, fill = "#0072B2", color = "white", alpha = 0.8) +
geom_vline(xintercept = c(6, 12), linetype = "dashed", color = "#D55E00") +
annotate("text",
x = 12.5, y = Inf, label = "12-month\n(annual)",
hjust = 0, vjust = 1.5, color = "#D55E00", fontface = "bold"
) +
annotate("text",
x = 6.5, y = Inf, label = "6-month\n(semi-annual)",
hjust = 0, vjust = 1.5, color = "#D55E00", fontface = "bold"
) +
labs(
x = "Dominant period (months)",
y = "Number of indicators",
title = "Distribution of dominant periods among periodic HMIS indicators"
) +
theme_minimal(base_size = 12) +
theme(
plot.title = element_text(face = "bold"),
plot.title.position = "plot",
panel.grid.minor = element_blank()
)
Strongest periodic signals
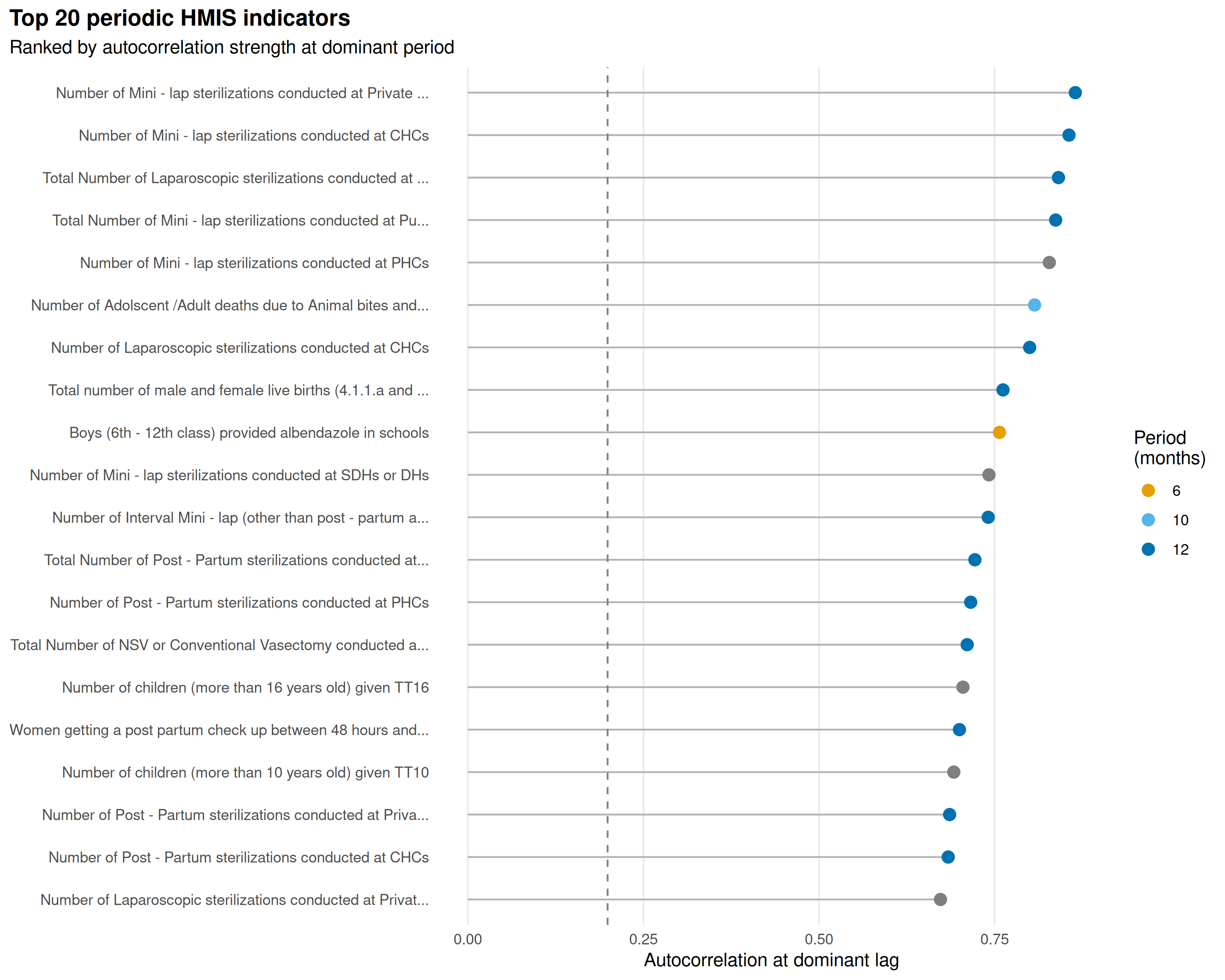
Top 20 indicators by autocorrelation at their dominant period:
top_periodic <- periodic |>
arrange(desc(acf_peak)) |>
head(20) |>
mutate(
short_name = ifelse(nchar(canonical_name) > 60,
paste0(substr(canonical_name, 1, 57), "..."),
canonical_name
),
short_name = make.unique(short_name),
short_name = factor(short_name, levels = rev(short_name))
)
ggplot(top_periodic, aes(acf_peak, short_name)) +
geom_segment(aes(xend = 0, yend = short_name), color = "grey70") +
geom_point(aes(color = factor(round(dominant_period))), size = 3) +
geom_vline(
xintercept = scan_results$acf_threshold[1],
linetype = "dashed", color = "grey50"
) +
scale_color_manual(
values = c(
"6" = "#E69F00", "12" = "#0072B2", "3" = "#009E73",
"4" = "#CC79A7", "9" = "#D55E00", "10" = "#56B4E9",
"5" = "#F0E442", "8" = "#999999", "7" = "#D55E00"
),
name = "Period\n(months)"
) +
labs(
x = "Autocorrelation at dominant lag",
y = NULL,
title = "Top 20 periodic HMIS indicators",
subtitle = "Ranked by autocorrelation strength at dominant period"
) +
theme_minimal(base_size = 11) +
theme(
plot.title = element_text(face = "bold"),
plot.title.position = "plot",
panel.grid.minor = element_blank(),
panel.grid.major.y = element_blank()
)
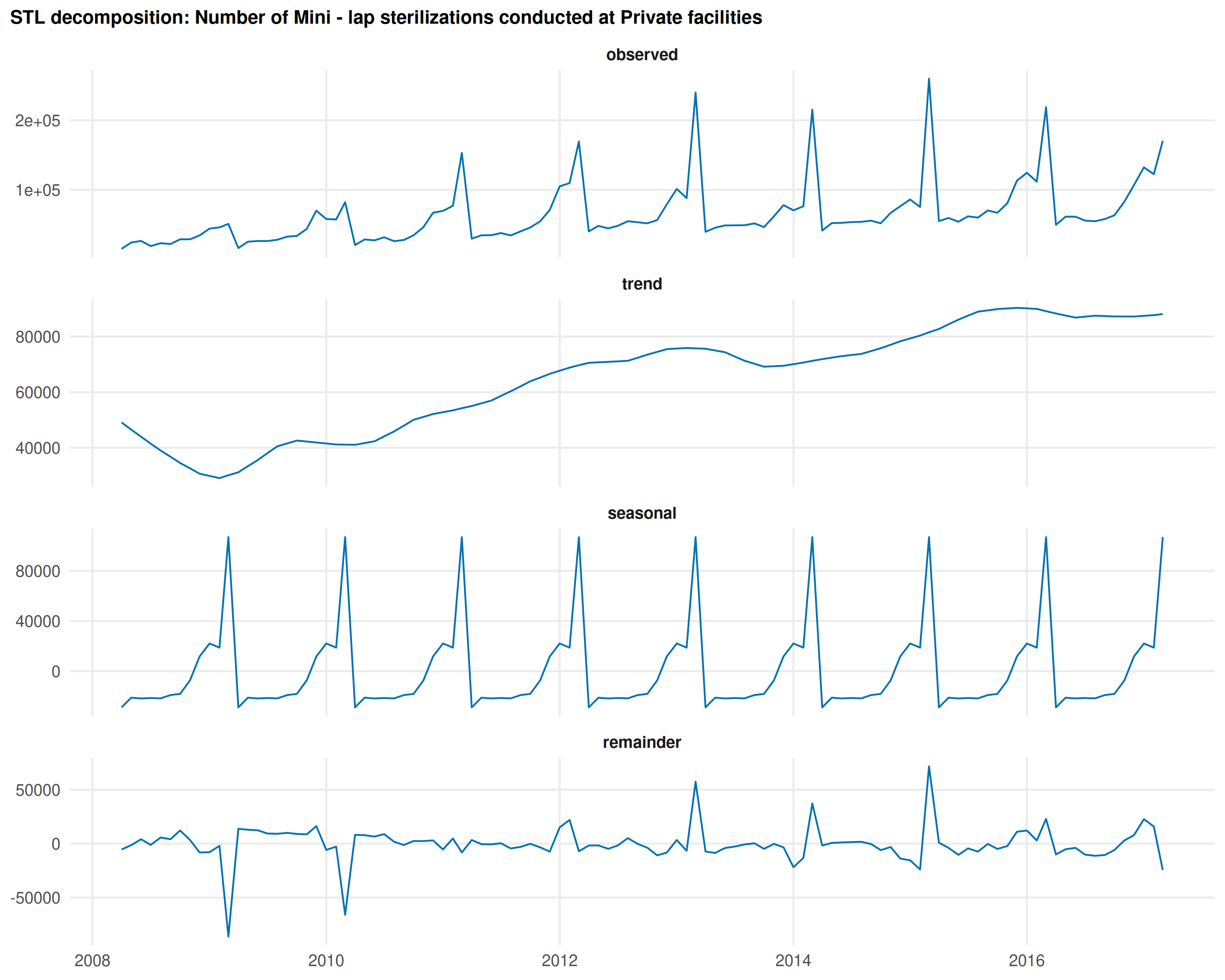
Decomposing the most seasonal indicator
top_name <- as.character(top_periodic$canonical_name[1])
national <- national_all |>
filter(canonical_name == top_name)
decomp <- decompose_series(national)
decomp_long <- decomp |>
pivot_longer(
cols = c(observed, trend, seasonal, remainder),
names_to = "component", values_to = "val"
) |>
mutate(component = factor(component,
levels = c("observed", "trend", "seasonal", "remainder")
))
ggplot(decomp_long, aes(date, val)) +
geom_line(color = "#0072B2", linewidth = 0.5) +
facet_wrap(~component, ncol = 1, scales = "free_y") +
labs(
x = NULL, y = NULL,
title = paste("STL decomposition:", top_name)
) +
theme_minimal(base_size = 12) +
theme(
strip.text = element_text(face = "bold"),
panel.grid.minor = element_blank(),
plot.title = element_text(face = "bold", size = 11),
plot.title.position = "plot"
)
All indicators: periodicity scores
All scanned indicators with periodicity results at the national level.
scan_results |>
arrange(desc(is_periodic), desc(acf_peak)) |>
dplyr::select(
Indicator = canonical_name,
Months = n_months,
Periodic = is_periodic,
`Period (months)` = dominant_period,
`Spectral power (%)` = spectral_power_pct,
`ACF peak` = acf_peak
) |>
DT::datatable(
filter = "top",
options = list(pageLength = 20, scrollX = TRUE),
rownames = FALSE
)