# Define non-state entities and UTs
exclude_states <- c(
"All India", "M/O Defence", "M/O Railways", "Chandigarh",
"A & N Islands", "Andaman & Nicobar Islands", "Lakshadweep",
"The Dadra And Nagar Haveli And Daman And Diu", "DNHDD",
"Dadra & Nagar Haveli and Daman & Diu",
"Jammu And Kashmir", "Jammu & Kashmir",
"Ladakh", "Puducherry", "Goa", "Telangana"
)
north_states <- c("Uttar Pradesh", "Bihar", "Punjab", "Haryana", "Rajasthan", "Madhya Pradesh", "Chandigarh")
north_eastern_states = c("Arunachal Pradesh", "Assam", "Manipur", "West Bengal", "Jharkhand", "Sikkim")
south_states <- c("Andhra Pradesh", "Telangana", "Karnataka", "Kerala", "Tamil Nadu")All cause-specific neonatal deaths
Indicator coverage
Cause Period Asphyxia 2008–2025 Sepsis 2008–2025 LBW 2008–2017 Prematurity 2023–2025 Others 2008–2025
Live births
births_old <- get_hmis(
"Total number of male and female live births",
sector = "Total",
to = "Mar 2017"
) |> add_calendar_vars()
births_new <- bind_rows(
get_hmis("Number of male live births", sector = "Total", from = "Apr 2017"),
get_hmis("Number of female live births", sector = "Total", from = "Apr 2017")
) |> add_calendar_vars()
b_collapsed <- bind_rows(births_old, births_new) |>
filter(!state %in% exclude_states) |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(births = sum(as.numeric(value), na.rm = TRUE), .groups = "drop")Asphyxia
# Asphyxia: Apr 2008–Mar 2017 (old, 24hr–4wk) + Apr 2017–Apr 2025 (new, 0–4wk)
nmr_asphyxia <- bind_rows(
get_hmis(
"Number of cases of Infant deaths 24 hours to 4 weeks of birth with the probable cause being Asphyxia",
sector = "Total",
category = "Total",
to = "Mar 2017"
),
get_hmis(
"Infant Deaths up to 4 weeks due to Asphyxia",
sector = "Total",
from = "Apr 2017"
)
) |>
add_calendar_vars() |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(deaths = sum(as.numeric(value), na.rm = TRUE), .groups = "drop") |>
inner_join(b_collapsed,
by = c("state", "monyear", "cal_year", "month", "month_num")) |>
mutate(nmr = (deaths / births) * 1000, cause = "Asphyxia")Sepsis
Neonatal sepsis, a systemic infection in the first 28 days of life, encompasses bloodstream infections, meningitis, and pneumonia.
# Sepsis: Apr 2008–Mar 2017 (old) + Apr 2017–Apr 2025 (new)
nmr_sepsis <- bind_rows(
get_hmis(
"Number of cases of Infant deaths 24 hours to 4 weeks of birth with the probable cause being Sepsis",
sector = "Total",
category = "Total",
to = "Mar 2017"
),
get_hmis(
"Infant Deaths up to 4 weeks due to Sepsis",
sector = "Total",
from = "Apr 2017"
)
) |>
add_calendar_vars() |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(deaths = sum(as.numeric(value), na.rm = TRUE), .groups = "drop") |>
inner_join(b_collapsed,
by = c("state", "monyear", "cal_year", "month", "month_num")) |>
mutate(nmr = (deaths / births) * 1000, cause = "Sepsis")LBW
# LBW: Apr 2008–Mar 2017 only (dropped from new series)
nmr_lbw <- get_hmis(
"Number of cases of Infant deaths 24 hours to 4 weeks of birth with the probable cause being LBW",
sector = "Total",
category = "Total",
to = "Mar 2017"
) |>
add_calendar_vars() |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(deaths = sum(as.numeric(value), na.rm = TRUE), .groups = "drop") |>
inner_join(b_collapsed,
by = c("state", "monyear", "cal_year", "month", "month_num")) |>
mutate(nmr = (deaths / births) * 1000, cause = "LBW")
# Other: Apr 2008–Mar 2017 (old) + Apr 2017–Apr 2025 (new)
nmr_other <- bind_rows(
get_hmis(
"Number of cases of Infant deaths 24 hours to 4 weeks of birth with the probable cause being other than Sepsis, Asphyxia and LBW",
sector = "Total",
category = "Total",
to = "Mar 2017"
),
get_hmis(
"Infant Deaths up to 4 weeks due to Other causes",
sector = "Total",
from = "Apr 2017"
)
) |>
add_calendar_vars() |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(deaths = sum(as.numeric(value), na.rm = TRUE), .groups = "drop") |>
inner_join(b_collapsed,
by = c("state", "monyear", "cal_year", "month", "month_num")) |>
mutate(nmr = (deaths / births) * 1000, cause = "Other")
# Prematurity: Apr 2023–Apr 2025 only
nmr_prematurity <- get_hmis(
"Neonatal Deaths up to 4 weeks due to complications of Prematurity",
sector = "Total",
from = "Apr 2023"
) |>
add_calendar_vars() |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(deaths = sum(as.numeric(value), na.rm = TRUE), .groups = "drop") |>
inner_join(b_collapsed,
by = c("state", "monyear", "cal_year", "month", "month_num")) |>
mutate(nmr = (deaths / births) * 1000, cause = "Prematurity")Ratio Attribution Calibration: Scaling Factor for Indicator Alignment
In April 2017, HMIS indicators for neonatal death causes transitioned from reporting deaths occurring between 24 hours and 4 weeks to including all deaths from 0 to 4 weeks.
Assumption: The proportion of 0-4 week deaths occurring in the first 24 hours is stable over time — specifically that the Apr 2017–Mar 2023 fraction applies to 2008–2017 data.
Calculate the State & Cause specific Multipliers
deaths_24hr_old <- get_hmis(
"Number of cases of Infant deaths within 24 hrs of birth",
sector = "Total",
to = "Mar 2017"
) |> add_calendar_vars()
# from Apr 2017
deaths_24hr_new <- get_hmis(
"Infant deaths within 24 hours (1 to 23 hours) of birth",
sector = "Total",
from = "Apr 2017",
to = "Mar 2023"
) |> add_calendar_vars()
# get total 24hr deaths (from Apr 2017, no cause breakdown)
deaths_24hr_summarised <- deaths_24hr_new |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(deaths_24hr = sum(as.numeric(value), na.rm = TRUE),
.groups = "drop")
# get total cause deaths (0-4 weeks)
# Total cause deaths post-2017 (denominator for frac_24hr)
deaths_cause_total <- bind_rows(
nmr_asphyxia |> filter(cal_year >= 2017,
!(cal_year == 2017 & month_num <= 3)),
nmr_sepsis |> filter(cal_year >= 2017,
!(cal_year == 2017 & month_num <= 3)),
nmr_other |> filter(cal_year >= 2017,
!(cal_year == 2017 & month_num <= 3))
) |>
group_by(state, monyear, cal_year, month, month_num) |>
summarise(deaths_all_causes = sum(deaths, na.rm = TRUE), .groups = "drop")
# frac_24hr: mean fraction of 0-4wk deaths occurring in first 24hrs
# Computed from Apr 2017–Mar 2023 where 24hr indicator exists
frac_24hr <- deaths_24hr_summarised |>
inner_join(deaths_cause_total,
by = c("state", "monyear", "cal_year", "month", "month_num")) |>
mutate(
frac = deaths_24hr / deaths_all_causes,
frac = pmin(pmax(frac, 0), 0.5)
) |>
group_by(state) |>
summarise(mean_frac_24hr = mean(frac, na.rm = TRUE), .groups = "drop")
# Apply calibration: scale up old series by 1/(1-frac_24hr)
all_causes_nmr_calibrated <- all_causes_nmr |>
left_join(frac_24hr, by = "state") |>
mutate(
scale_factor = 1 / (1 - coalesce(mean_frac_24hr, 0.15)),
# 0.15 fallback for states with no post-2017 24hr data
nmr_adj = if_else(
(cal_year < 2017 | (cal_year == 2017 & month_num <= 3)) &
cause %in% c("Asphyxia", "Sepsis", "Other"),
nmr * scale_factor,
nmr
)
)Seasonality across months | Normalized heatmaps
Min‑max normalization: highlights seasonal peaks relative to each state’s own range.
# Min-Max normalize calibrated NMR per state and cause
nmr_minmax_adj <- all_causes_nmr_calibrated |>
# compute median adjusted NMR per state x cause x month across all years
group_by(state, cause, month) |>
summarise(med_nmr = median(nmr_adj, na.rm = TRUE), .groups = "drop") |>
mutate(month = factor(month, levels = month.abb)) |>
# normalize within each state x cause
group_by(state, cause) |>
mutate(
s_min = min(med_nmr, na.rm = TRUE),
s_max = max(med_nmr, na.rm = TRUE),
nmr_norm = if_else(
s_max > s_min,
(med_nmr - s_min) / (s_max - s_min),
0.5 # flat states -> mid-range
)
) |>
ungroup()
# Updated state ordering function for adjusted data
order_states_by_peak_adj <- function(df, cause_filter) {
df |>
filter(cause == cause_filter) |>
group_by(state) |>
slice_max(nmr_norm, n = 1, with_ties = FALSE) |>
arrange(as.integer(month)) |>
pull(state)
}
# Heatmap function for adjusted data
make_minmax_heatmap_adj <- function(cause_filter, title_label) {
state_order <- order_states_by_peak_adj(nmr_minmax_adj, cause_filter)
nmr_minmax_adj |>
filter(cause == cause_filter) |>
mutate(state = factor(state, levels = state_order)) |>
ggplot(aes(x = month, y = state, fill = nmr_norm)) +
geom_tile(color = "white", linewidth = 0.2) +
scale_fill_distiller(
palette = "YlOrBr",
direction = 1,
name = "Seasonal\nindex",
limits = c(0, 1)
) +
labs(
title = paste0("Min-Max Normalized Adjusted NMR: ", title_label),
caption = "States ordered by peak month",
x = NULL, y = NULL
) +
theme_hmis() +
theme(
panel.grid = element_blank(),
axis.text.x = element_text(hjust = 1),
axis.text = element_text(face = "bold"),
legend.position = "top"
) +
guides(
fill = guide_colorbar(
barwidth = 12,
barheight = 0.55
)
) +
labs_hmis()
}
make_minmax_heatmap_adj("Asphyxia", "Asphyxia")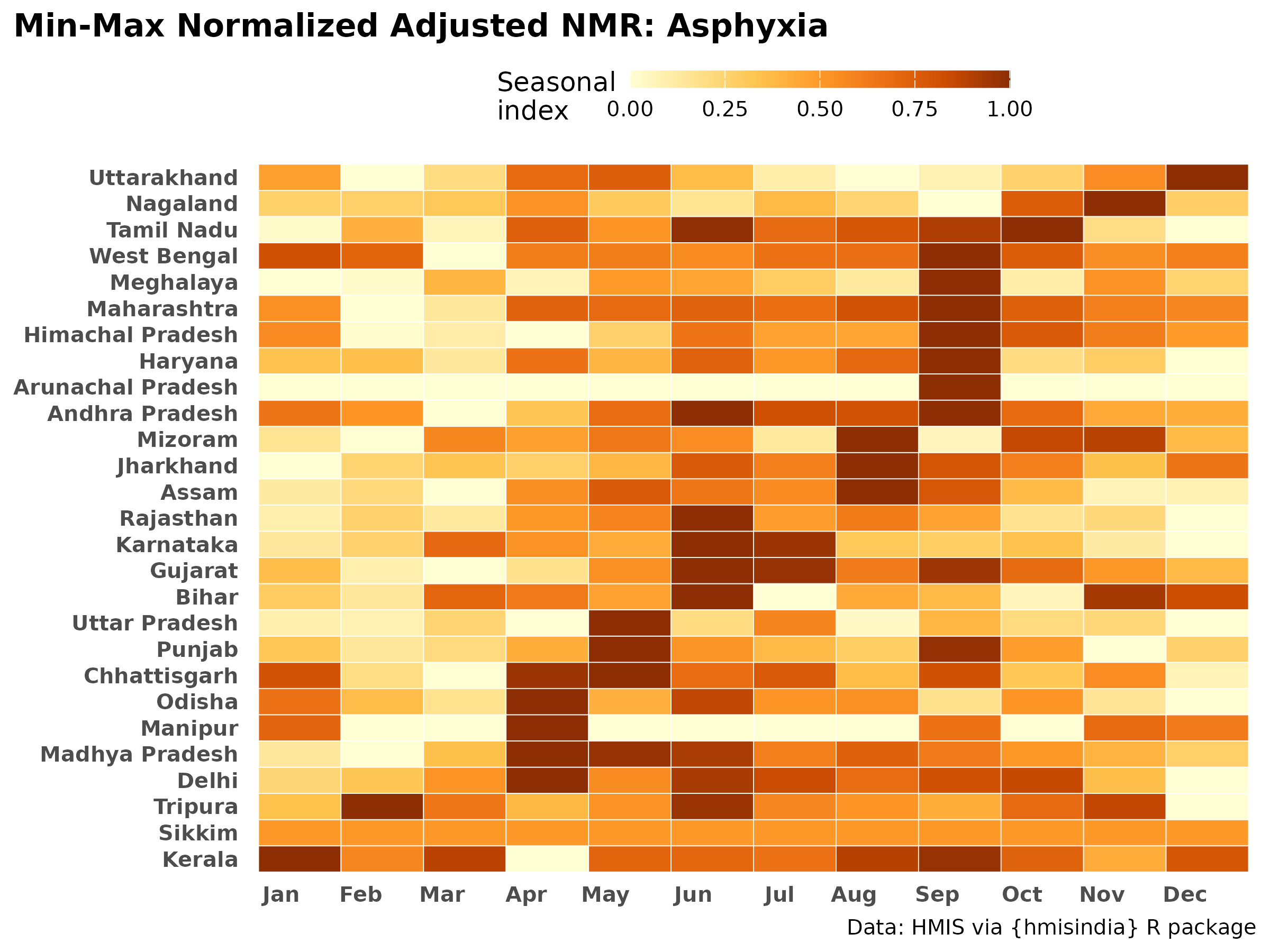
make_minmax_heatmap_adj("Sepsis", "Sepsis")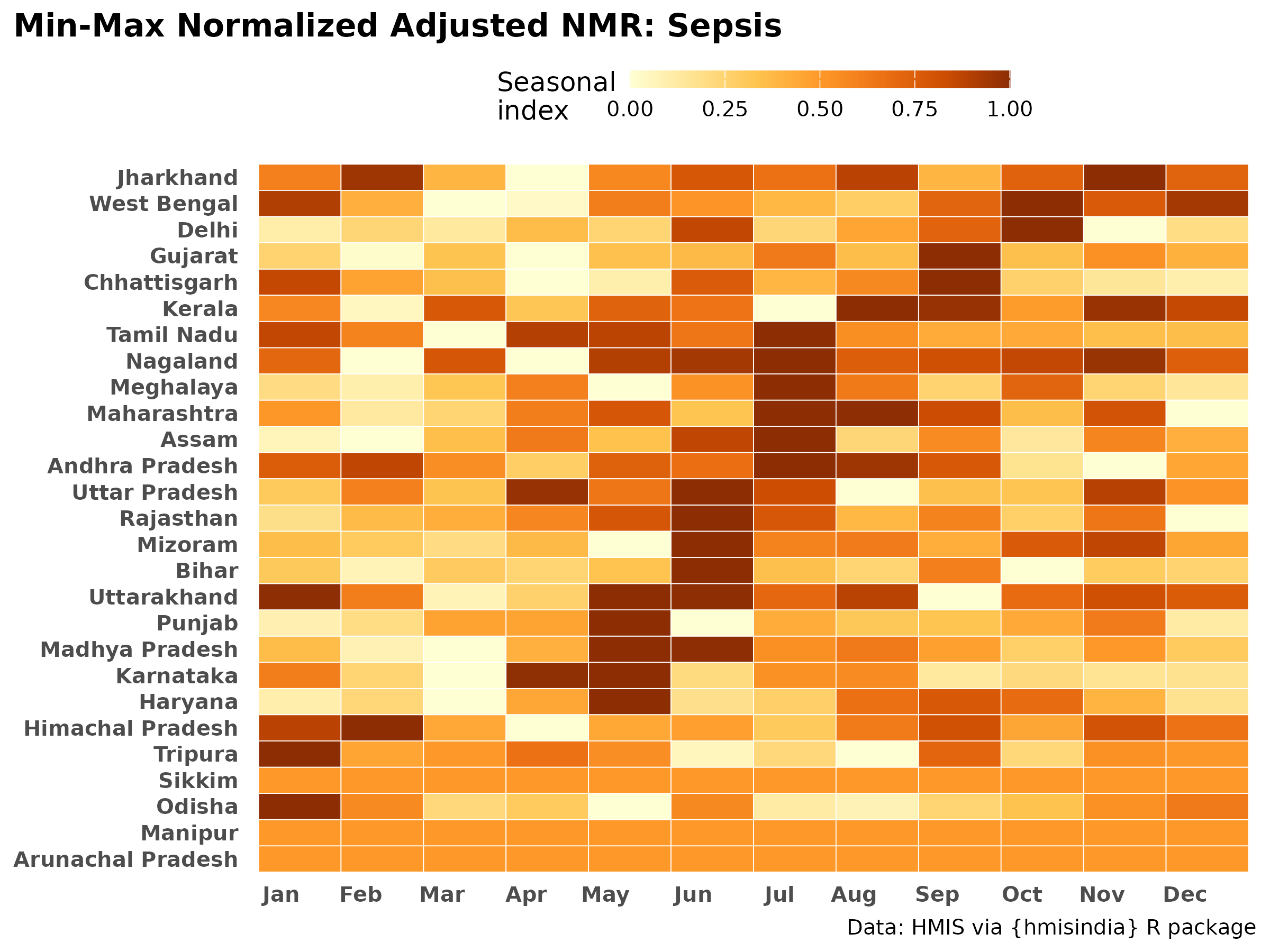
Z‑score normalization: highlights months unusually high/low compared to that state’s average
# Z-score normalize
nmr_zscore_adj <- all_causes_nmr_calibrated |>
# Normalize within year first
group_by(state, cause, cal_year) |>
mutate(
med_val = median(nmr_adj, na.rm = TRUE),
mad_val = mad(nmr_adj, na.rm = TRUE),
z = if_else(mad_val == 0, 0,
(nmr_adj - med_val) / mad_val)
) |>
ungroup() |>
# Cap at ±3
mutate(z = pmin(pmax(z, -3), 3)) |>
group_by(state, cause, month) |>
summarise(med_z = median(z, na.rm = TRUE), .groups = "drop") |>
mutate(month = factor(month, levels = month.abb))
# Asphyxia
ggplot(filter(nmr_zscore_adj, cause == "Asphyxia"),
aes(x = month, y = state, fill = med_z)) +
geom_tile(color = "white", linewidth = 0.1) +
scale_fill_distiller(
palette = "PuOr",
direction = 1
) +
labs(title = "Z-score Normalized NMR: Asphyxia",
x = NULL, y = NULL, fill = "Z-score") +
theme_minimal() +
theme(axis.text.x = element_text(hjust = 1),
legend.position = "top"
) +
guides(
fill = guide_colorbar(
barwidth = 12,
barheight = 0.55
)
) +
labs_hmis()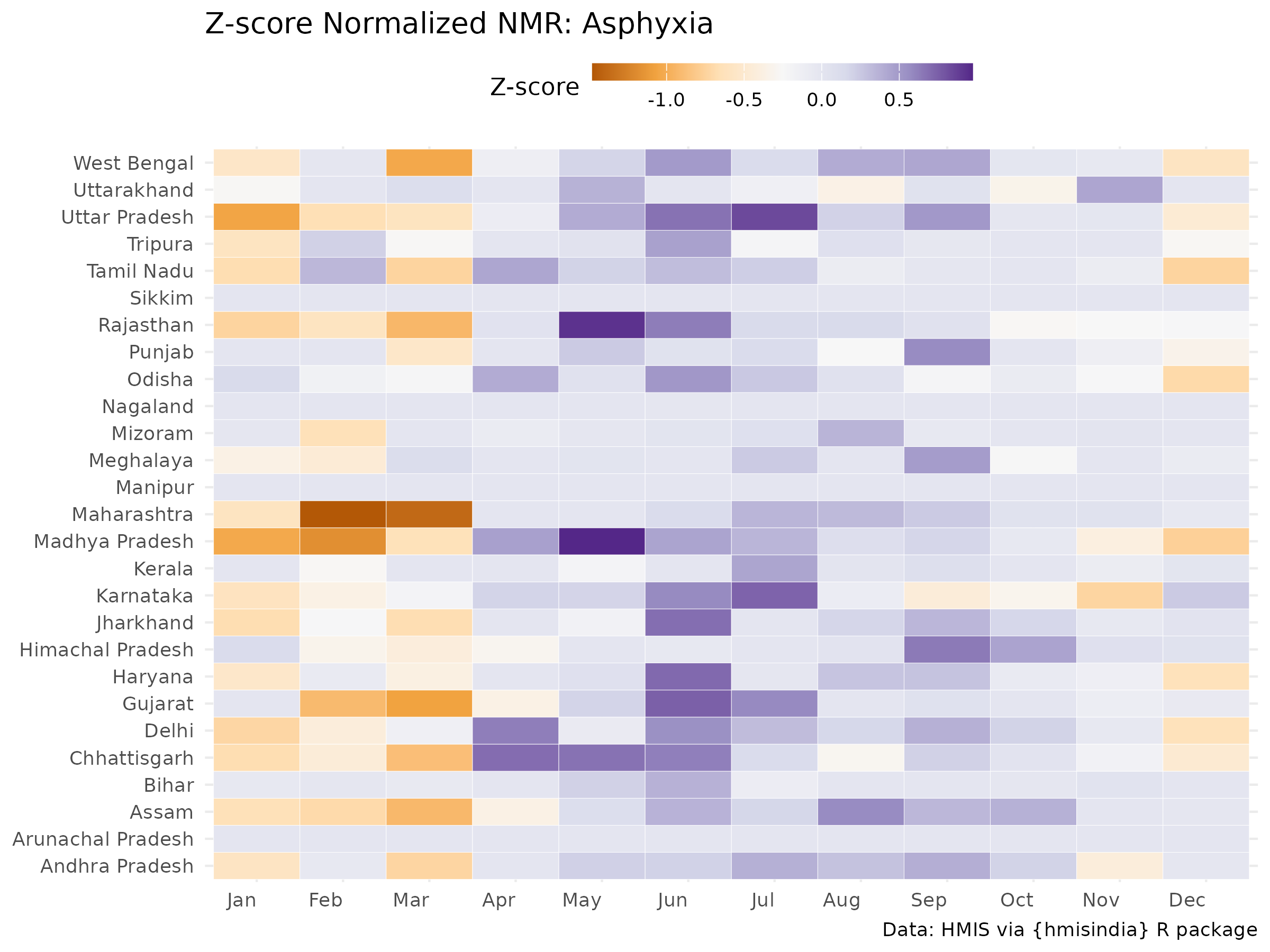
# Sepsis
ggplot(filter(nmr_zscore_adj, cause == "Sepsis"),
aes(x = month, y = state, fill = med_z)) +
geom_tile(color = "white", linewidth = 0.1) +
scale_fill_distiller(
palette = "PuOr",
direction = 1
) +
labs(title = "Z-score Normalized Adjusted NMR: Sepsis",
#subtitle = " ",
x = NULL, y = NULL, fill = "z-score") +
guides(
fill = guide_colorbar(
barwidth = 12,
barheight = 0.55
)
) +
theme_hmis() +
theme(axis.text = element_text(face = "bold"),
axis.text.x = element_text(hjust = 1),
legend.position = "top") +
labs_hmis()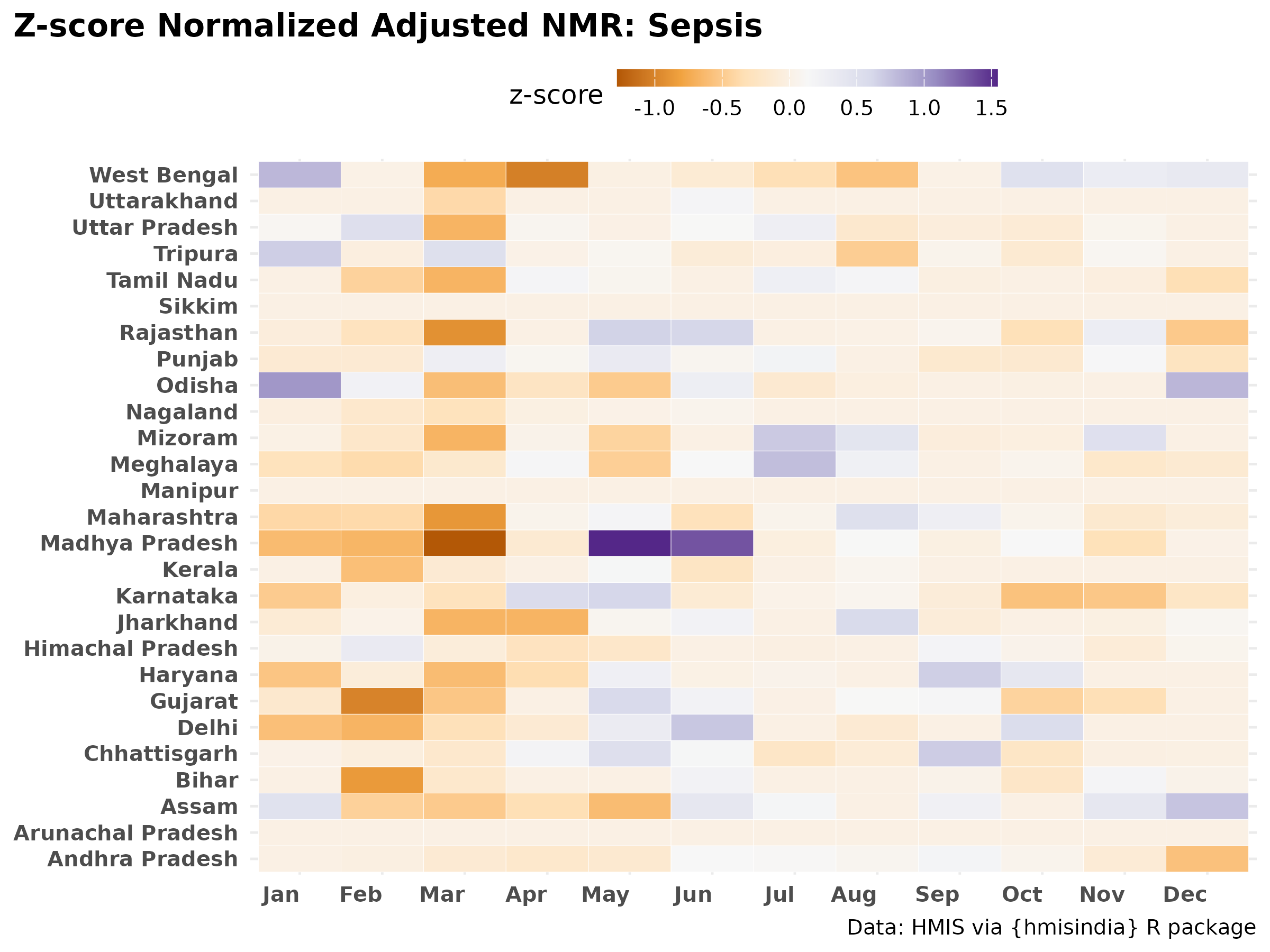 ## Is Asphyxia death correlated with Sepsis?
## Is Asphyxia death correlated with Sepsis?
# Monthly median for Calibrated Asphyxia and Sepsis
cause_month_adj <- all_causes_nmr_calibrated %>%
filter(cause %in% c("Asphyxia", "Sepsis")) %>%
group_by(month, cal_year, cause) %>%
summarise(median_nmr_adj = median(nmr_adj, na.rm = TRUE), .groups = "drop") %>%
pivot_wider(names_from = cause, values_from = median_nmr_adj)
# Calibrated Correlation coefficient
cor_val_adj <- cor(cause_month_adj$Asphyxia, cause_month_adj$Sepsis, use = "complete.obs")
print(paste("Standard Correlation (Calibrated):", round(cor_val_adj, 3)))
#> [1] "Standard Correlation (Calibrated): 0.796"
#Current cor() on monthly medians pooled across all years.
#A partial correlation controlling for month would be more informative.
cor_data_adj <- cause_month_adj %>%
mutate(month_num = match(month, month.abb)) %>%
filter(!is.na(Asphyxia) & !is.na(Sepsis))
partial_res_adj <- pcor.test(cor_data_adj$Asphyxia, cor_data_adj$Sepsis, cor_data_adj$month_num)
print(paste("Partial Correlation (Calibrated):", round(partial_res_adj$estimate, 3)))
#> [1] "Partial Correlation (Calibrated): 0.795"The partial correlation barely changed after controlling for month. This means the asphyxia-sepsis correlation is not just because both peak in monsoon together, but there might be state-level co-occurrence. States with high asphyxia tend to also have high sepsis regardless of month. This points to a common structural driver — likely healthcare system quality or SNCU capacity rather than a shared environmental exposure.
# Calibrated Monthly National Averages
seasonal_corr_adj <- all_causes_nmr_calibrated %>%
filter(cause %in% c("Asphyxia", "Sepsis")) %>%
group_by(month, cause) %>%
summarise(med_nmr_adj = median(nmr_adj, na.rm = TRUE), .groups = "drop") %>%
mutate(month = factor(month, levels = month.abb)) %>%
arrange(cause, month) %>%
group_by(cause) %>%
mutate(rel_change_adj = (med_nmr_adj - lag(med_nmr_adj)) / lag(med_nmr_adj) * 100) %>%
ungroup()
ggplot(seasonal_corr_adj, aes(x = month, y = rel_change_adj, group = cause, color = cause)) +
geom_line(linewidth = 1) +
geom_hline(yintercept = 0, linetype = "dotted", color = "grey40") +
theme_minimal() +
scale_color_manual(values = c(
"Asphyxia" = "steelblue",
"Sepsis" = "tan3"
)) +
labs(
title = "Calibrated Monthly Growth Rate: Asphyxia vs Sepsis",
subtitle = "% Relative change in Adjusted NMR compared to previous month",
y = "% Change from Previous Month",
x = NULL) +
theme_hmis() +
theme(legend.position = "top",
legend.title = element_blank(),
axis.text = element_text(face = "bold")) +
labs_hmis()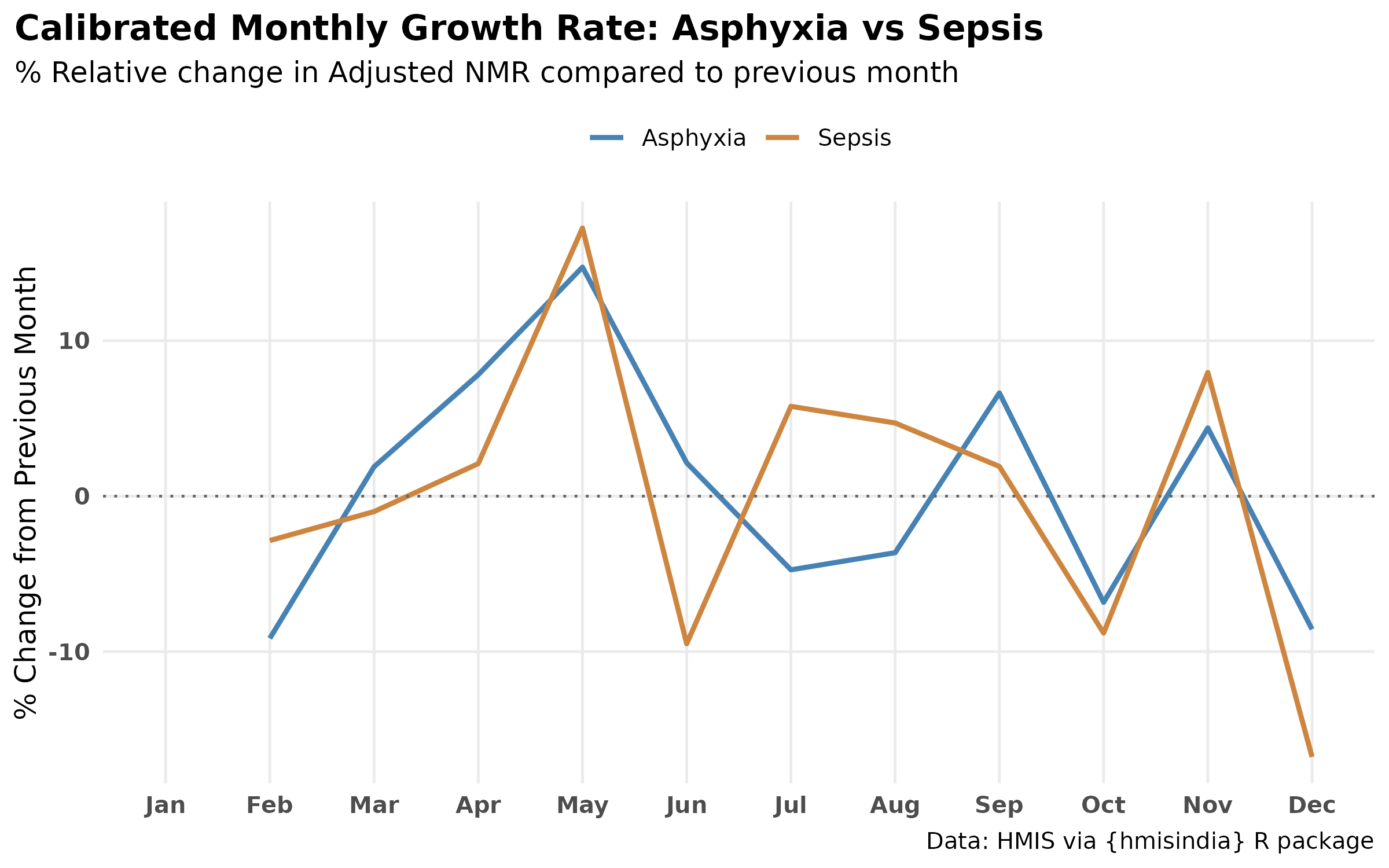
# Monthly averages for Calibrated Asphyxia and Sepsis
cause_month_adj_avg <- all_causes_nmr_calibrated %>%
filter(cause %in% c("Asphyxia", "Sepsis")) %>%
group_by(month, cal_year, cause) %>%
summarise(avg_nmr_adj = mean(nmr_adj, na.rm = TRUE), .groups = "drop") %>%
pivot_wider(names_from = cause, values_from = avg_nmr_adj)
# Calibrated Correlation coefficient (using Averages)
cor_val_adj_avg <- cor(cause_month_adj_avg$Asphyxia,
cause_month_adj_avg$Sepsis,
use = "complete.obs")
# % Relative change across month (National Averages)
seasonal_corr_adj_avg <- all_causes_nmr_calibrated %>%
filter(cause %in% c("Asphyxia", "Sepsis")) %>%
group_by(month, cause) %>%
summarise(avg_nmr_adj = mean(nmr_adj, na.rm = TRUE), .groups = "drop") %>%
mutate(month = factor(month, levels = month.abb)) %>%
arrange(cause, month) %>%
group_by(cause) %>%
mutate(rel_change = (avg_nmr_adj - lag(avg_nmr_adj)) / lag(avg_nmr_adj) * 100) %>%
ungroup() %>%
filter(!is.na(rel_change))
# Plotting Calibrated Growth Rate
ggplot(seasonal_corr_adj_avg, aes(x = month, y = rel_change, group = cause, color = cause)) +
geom_line(linewidth = 1) +
geom_hline(yintercept = 0, linetype = "dotted", color = "grey40") +
theme_minimal() +
scale_color_manual(values = c("Asphyxia" = "steelblue4", "Sepsis" = "tan3")) +
labs(
title = "Calibrated Monthly Growth Rate: Asphyxia vs Sepsis",
subtitle = "% Relative change in Adjusted NMR (National Average)",
y = "% Change from Previous Month",
x = NULL
) +
theme_hmis() +
theme(legend.position = "top",
legend.title = element_blank(),
axis.text = element_text(face = "bold")) +
labs_hmis()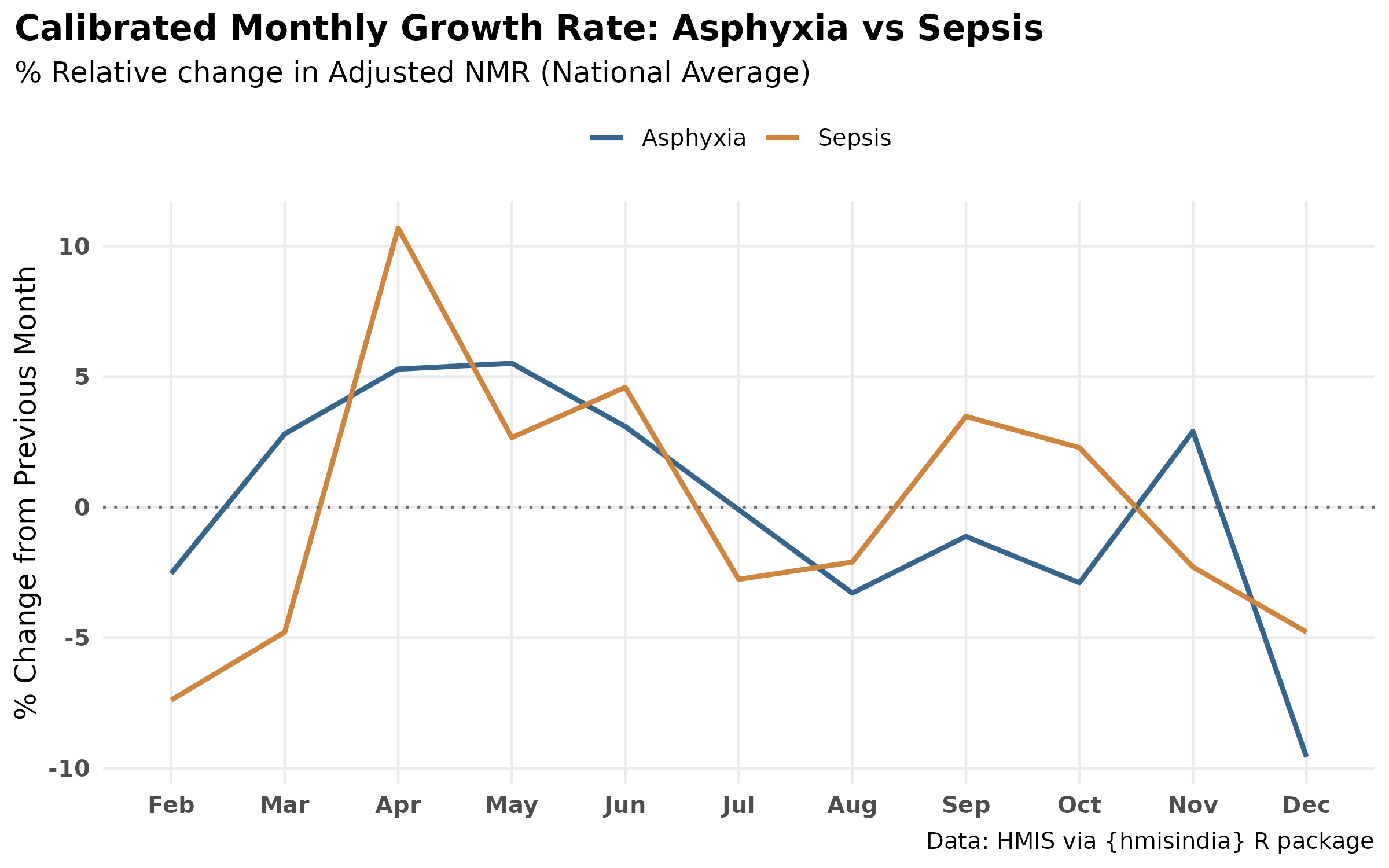
# National trend using calibrated NMR
national_trend_adj <- all_causes_nmr_calibrated %>%
filter(cause %in% c("Asphyxia", "Sepsis", "LBW", "Other", "Prematurity")) %>%
group_by(cal_year, cause) %>%
summarise(national_med = median(nmr_adj, na.rm = TRUE), .groups = "drop")
ggplot(national_trend_adj, aes(x = cal_year, y = national_med, color = cause)) +
geom_line(linewidth = 1) +
geom_vline(xintercept = 2023, linetype = "dashed",
color = "#E69F00", linewidth = 0.6) +
annotate("text", x = 2023.1, y = Inf, vjust = 1.5, hjust = 0,
label = "Prematurity\nadded", size = 2.8, color = "#E69F00") +
geom_vline(xintercept = 2017, linetype = "dotted",
color = "grey40", linewidth = 0.7) +
annotate(geom = "text", x = 2015, y = 1.4,
hjust = 0, size = 3, color = "grey4",
label = "Apr 2017 indicator\nrename") +
scale_color_manual(values = c(
"Asphyxia" = "steelblue",
"Sepsis" = "tan3",
"LBW" = "#009E73",
"Other" = "#999999",
"Prematurity" = "#E69F00"
)) +
labs(
title = "National NMR Trend",
#subtitle = " ",
y = "Adjusted NMR",
x = NULL
) +
theme_hmis() +
theme(legend.position = "top",
legend.title = element_blank(),
axis.text = element_text(face = "bold")) +
labs_hmis()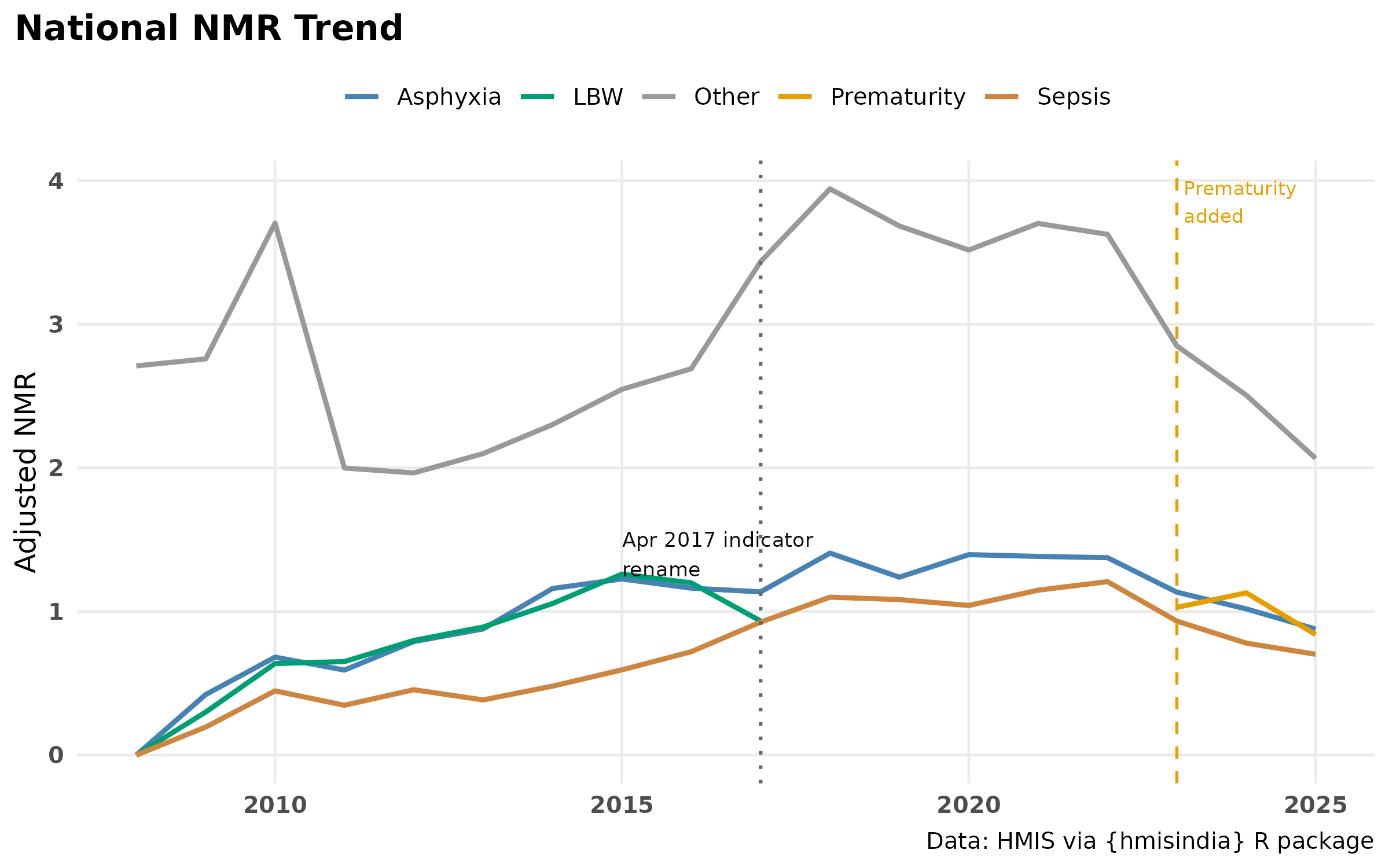
# lines starting from 0 in 2008, because the first year has incomplete data (Apr 2008 = start mid-year). Maybe filter out 2008
# State-level summary using calibrated NMR
cause_state_adj <- all_causes_nmr_calibrated |>
group_by(state, cal_year, cause) |>
summarise(avg_nmr_adj = mean(nmr_adj, na.rm = TRUE), .groups = "drop")
# Define high-burden list
high_burden <- c("Uttar Pradesh", "Bihar", "Assam", "West Bengal",
"Madhya Pradesh", "Chhattisgarh", "Jharkhand",
"Odisha", "Rajasthan")
# Faceted Asphyxia Trend
ggplot(filter(cause_state_adj, cause == "Asphyxia", state %in% high_burden),
aes(x = cal_year, y = avg_nmr_adj)) +
geom_line(color = "steelblue", linewidth = 0.6) +
#geom_smooth(method = "loess", span = 0.2, color = "steelblue", se = FALSE) +
facet_wrap(~state, scales = "free_y") +
labs(
title = "Asphyxia NMR: High-Burden States",
#subtitle = "",
y = "Adjusted NMR (per 1,000 live births)",
x = NULL
) +
theme_hmis() +
labs_hmis()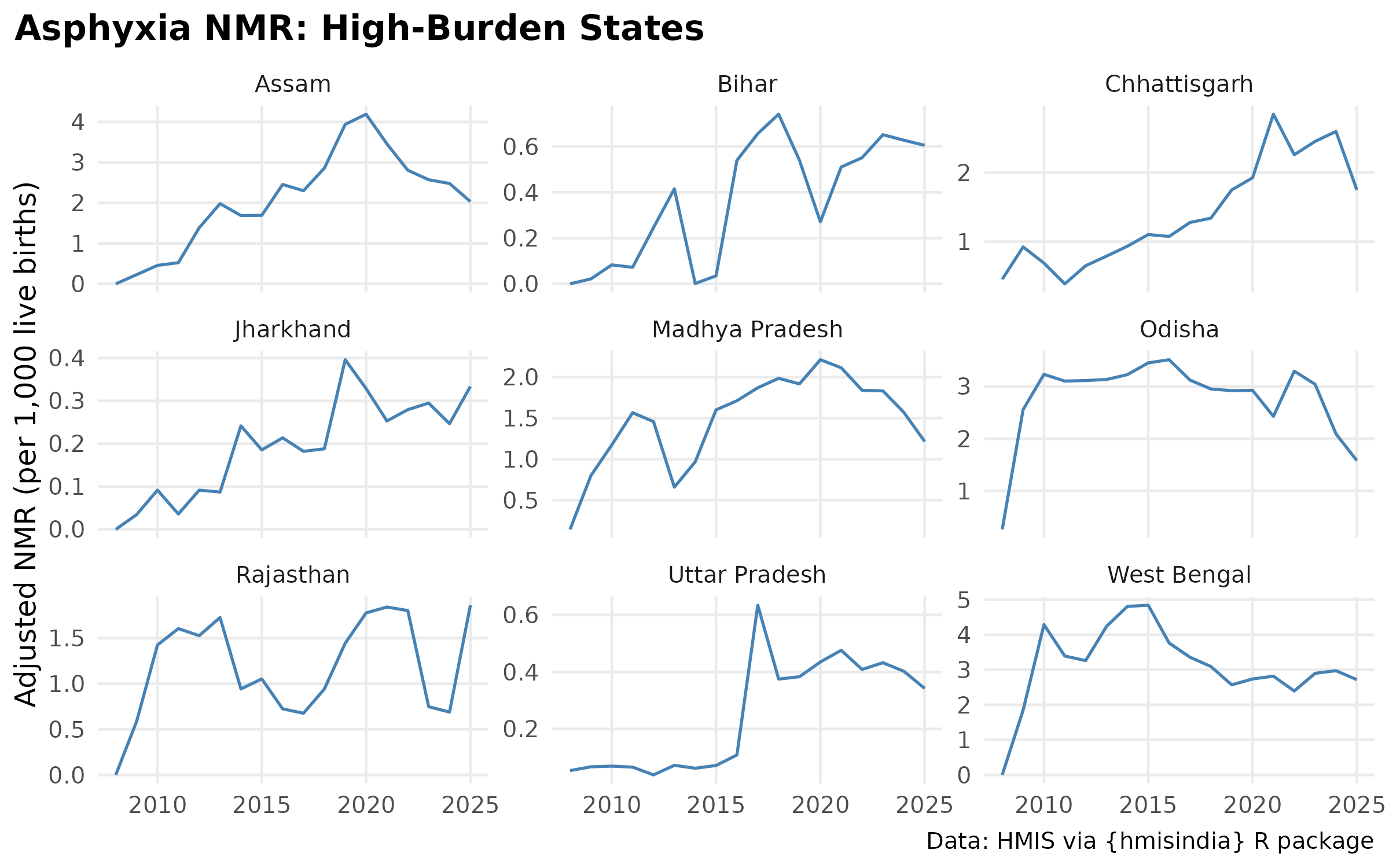
# Faceted Sepsis Trend
ggplot(filter(cause_state_adj, cause == "Sepsis", state %in% high_burden),
aes(x = cal_year, y = avg_nmr_adj)) +
geom_line(color = "tan3", linewidth = 0.6) +
facet_wrap(~state, scales = "free_y") +
labs(
title = "Sepsis NMR: High-Burden States",
#subtitle = "",
y = "Adjusted NMR (per 1,000 live births)",
x = NULL
) +
theme_hmis() +
labs_hmis()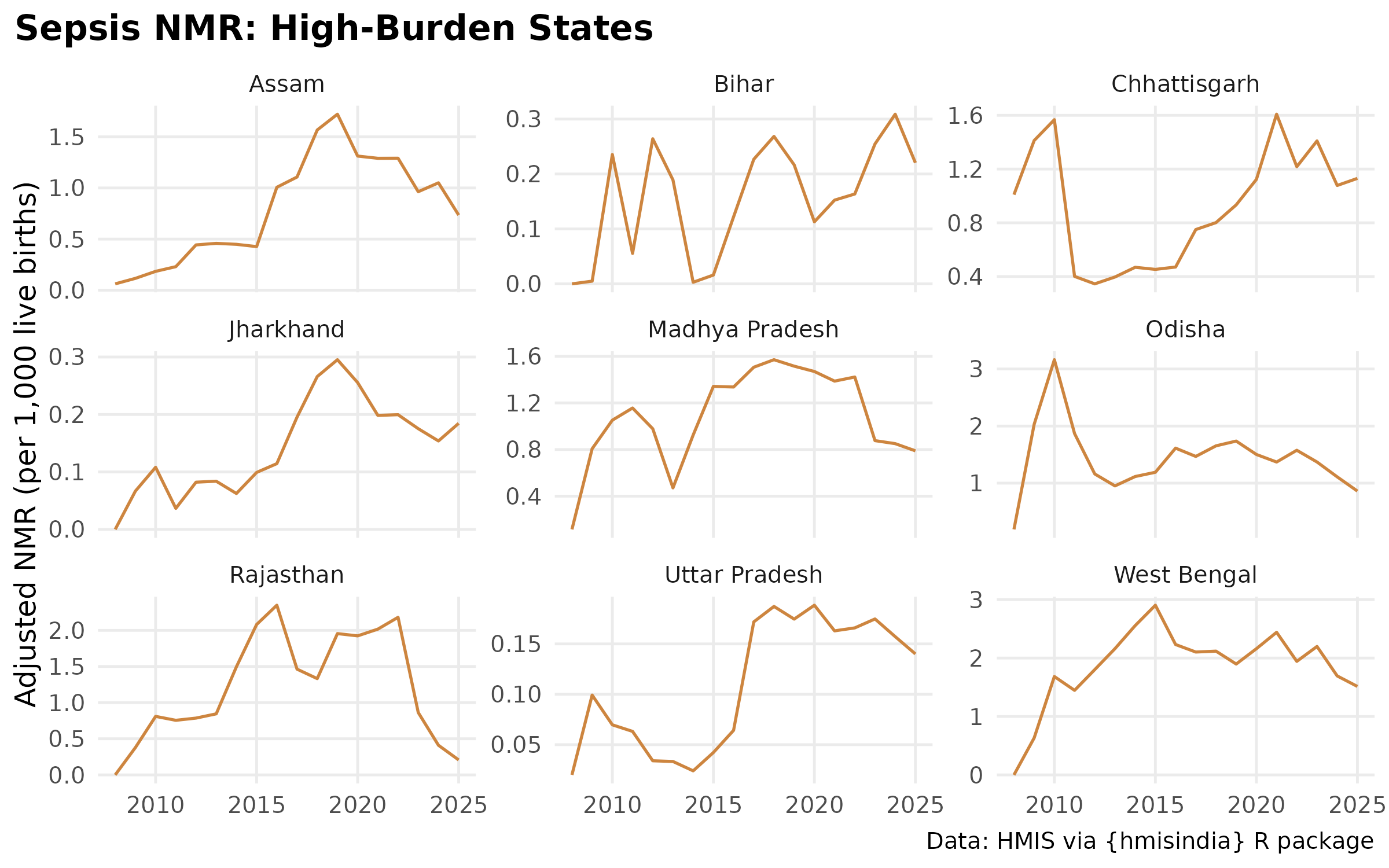
# lines starting from 0 in 2008, because the first year has incomplete data (Apr 2008 = start mid-year). Maybe filter out 2008
region_map <- data.frame(
state = c(
# North India
"Uttar Pradesh", "Bihar", "Punjab", "Haryana", "Rajasthan", "Madhya Pradesh", "Chandigarh", "Uttarakhand", "Himachal Pradesh",
# North Eastern India
"Arunachal Pradesh", "Assam", "Manipur", "West Bengal", "Jharkhand", "Sikkim", "Meghalaya", "Mizoram", "Nagaland", "Tripura",
# South India
"Andhra Pradesh", "Telangana", "Karnataka", "Kerala", "Tamil Nadu",
# West & Central
"Maharashtra", "Gujarat", "Chhattisgarh", "Odisha"
),
region = c(
rep("North India", 9),
rep("North East", 10),
rep("South India", 5),
rep("West/Central", 4)
)
)
scatter_data_adj <- all_causes_nmr_calibrated |>
filter(cause %in% c("Asphyxia", "Sepsis")) |>
# Median adjusted NMR per state x month x cause
group_by(state, month, cause) |>
summarise(med_nmr_adj = median(nmr_adj, na.rm = TRUE), .groups = "drop") |>
pivot_wider(names_from = cause, values_from = med_nmr_adj) |>
left_join(region_map, by = "state") |>
filter(!is.na(region), !is.na(Asphyxia), !is.na(Sepsis)) |>
mutate(month = factor(month, levels = month.abb))
region_colours <- c(
"North India" = "#D55E00",
"South India" = "steelblue",
"North East" = "#009E73",
"West/Central" = "#CC79A7"
)
ggplot(scatter_data_adj, aes(x = Asphyxia, y = Sepsis, color = region)) +
coord_cartesian(xlim = c(0, 3.5), ylim = c(0, 2.5)) +
geom_point(alpha = 0.5, size = 2.5) +
geom_smooth(method = "lm", se = FALSE,
color = "grey30", linewidth = 0.7, linetype = "dotted") +
geom_abline(slope = 1, intercept = 0,
linetype = "dashed", color = "grey70", linewidth = 0.5) +
annotate("text", x = 0.3, y = 2.3, hjust = 0, size = 3.5,
label = "r = 0.80 | partial r = 0.80\n(controlling for month)") +
geom_text_repel(
data = scatter_data_adj |>
group_by(state) |>
summarise(Asphyxia = median(Asphyxia), Sepsis = median(Sepsis)) |>
left_join(region_map, by = "state"),
aes(label = state),
size = 3, fontface = "bold", max.overlaps = 15
) +
scale_color_manual(values = region_colours, name = NULL) +
labs(
title = "Asphyxia vs Sepsis NMR — State × Month combinations",
x = "Asphyxia NMR (per 1,000 live births)",
y = "Sepsis NMR (per 1,000 live births)"
) +
theme_hmis() +
theme(
plot.title.position = "plot",
panel.grid.minor = element_blank(),
legend.position = "top"
) +
labs_hmis()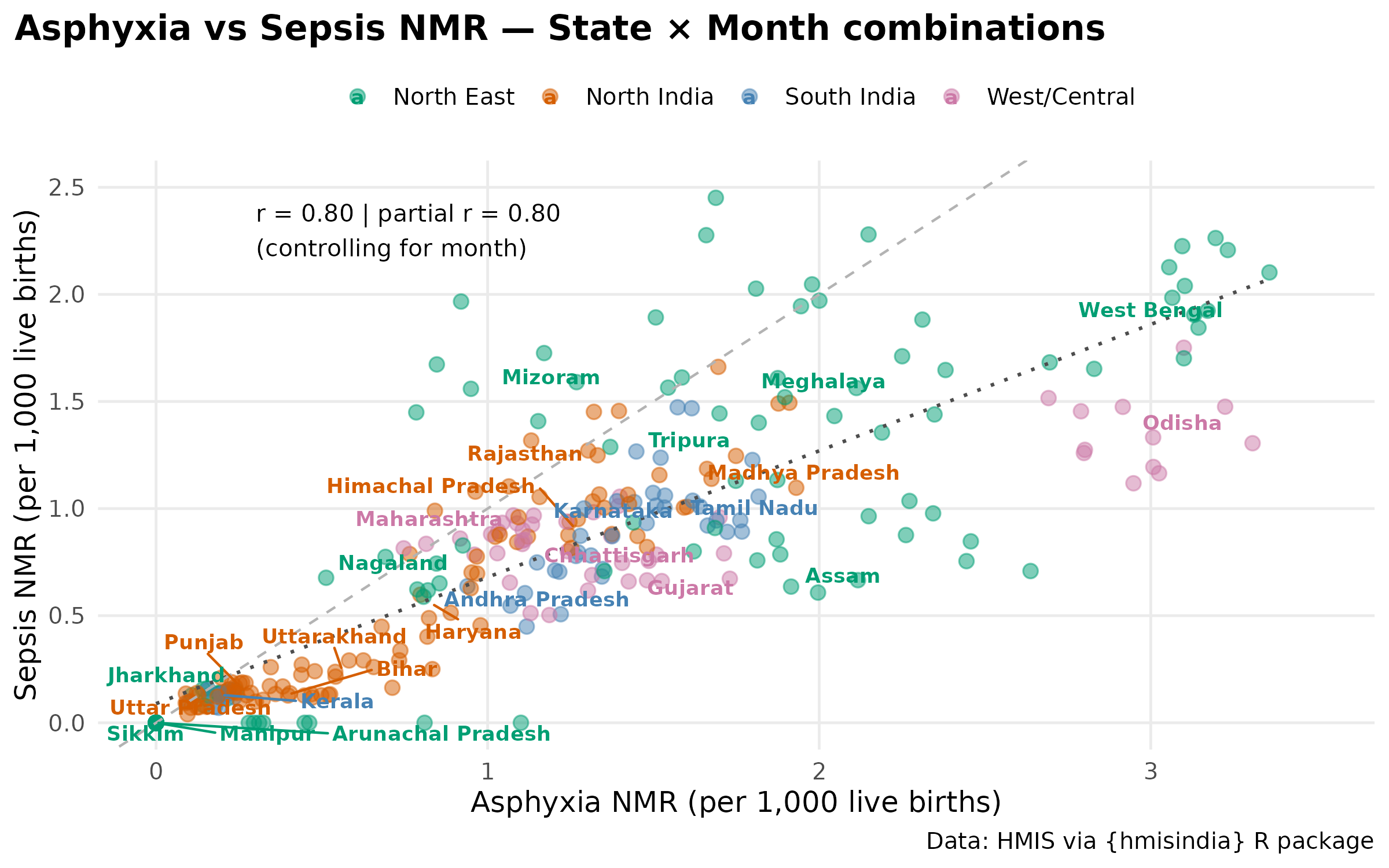
# Periodicity comparison: full series vs new series — by cause
causes_to_test <- c("Asphyxia", "Sepsis")
# Full series
periodicity_full <- all_causes_nmr_calibrated %>%
filter(cause %in% causes_to_test) %>%
group_by(monyear, cause) %>%
summarise(nmr_adj = median(nmr_adj, na.rm = TRUE), .groups = "drop")
# New series only
periodicity_new <- all_causes_nmr_calibrated %>%
filter(cause %in% causes_to_test,
cal_year >= 2018, cal_year <= 2024) %>%
group_by(monyear, cause) %>%
summarise(nmr_adj = median(nmr_adj, na.rm = TRUE), .groups = "drop")
# Run for both series and both causes
results <- list()
for (s in c("full", "new")) {
df <- if (s == "full") periodicity_full else periodicity_new
for (cause_name in causes_to_test) {
res <- detect_periodicity_ls(
df %>% filter(cause == cause_name),
value_col = "nmr_adj",
time_col = "monyear",
detrend = TRUE,
period_max = 24
)
results[[paste(cause_name, s, sep = "_")]] <- list(
cause = cause_name,
series = s,
periodic = res$is_periodic,
period = round(res$dominant_period, 1),
fap = signif(res$fap, 3),
peak_power = round(res$peak_power, 2),
result = res
)
}
}
# Summary table
results_df <- purrr::map_dfr(results, function(r) {
tibble(
cause = r$cause,
series = r$series,
periodic = r$periodic,
period = r$period,
fap = r$fap,
power = r$peak_power
)
}) |>
mutate(series = if_else(series == "full",
"Full (2008-2025)",
"New only (2018-2024)")) |>
arrange(cause, series)
print(results_df)
#> # A tibble: 4 × 6
#> cause series periodic period fap power
#> <chr> <chr> <lgl> <dbl> <dbl> <dbl>
#> 1 Asphyxia Full (2008-2025) FALSE 11.9 0.102 6.38
#> 2 Asphyxia New only (2018-2024) TRUE 11.8 0.0332 6.63
#> 3 Sepsis Full (2008-2025) FALSE 11.9 0.0992 6.41
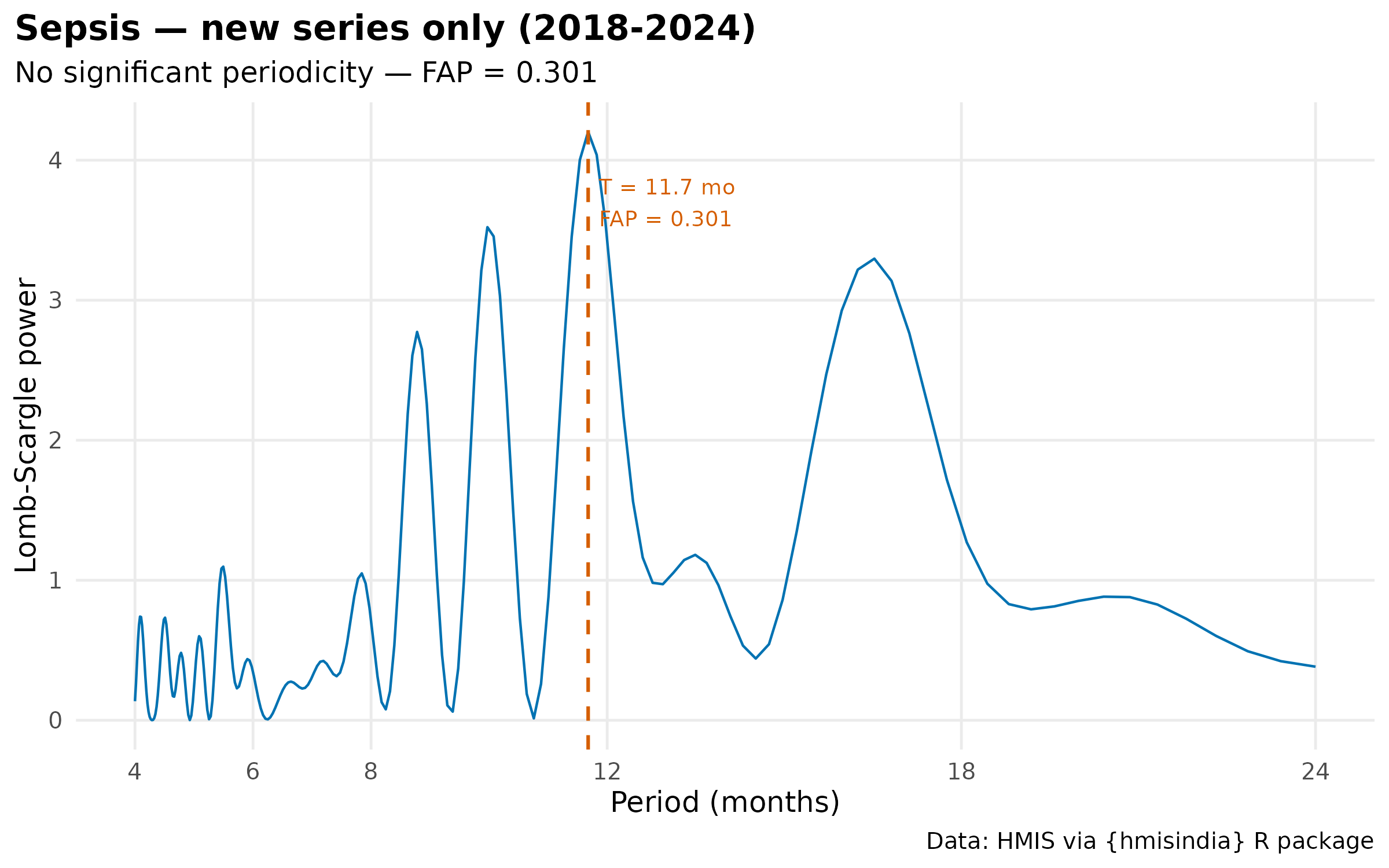
#> 4 Sepsis New only (2018-2024) FALSE 11.7 0.301 4.2
# Plot all four periodograms
plot(results[["Asphyxia_full"]]$result) +
labs(title = "Asphyxia — full series (2008-2025)") +
theme_hmis() + labs_hmis()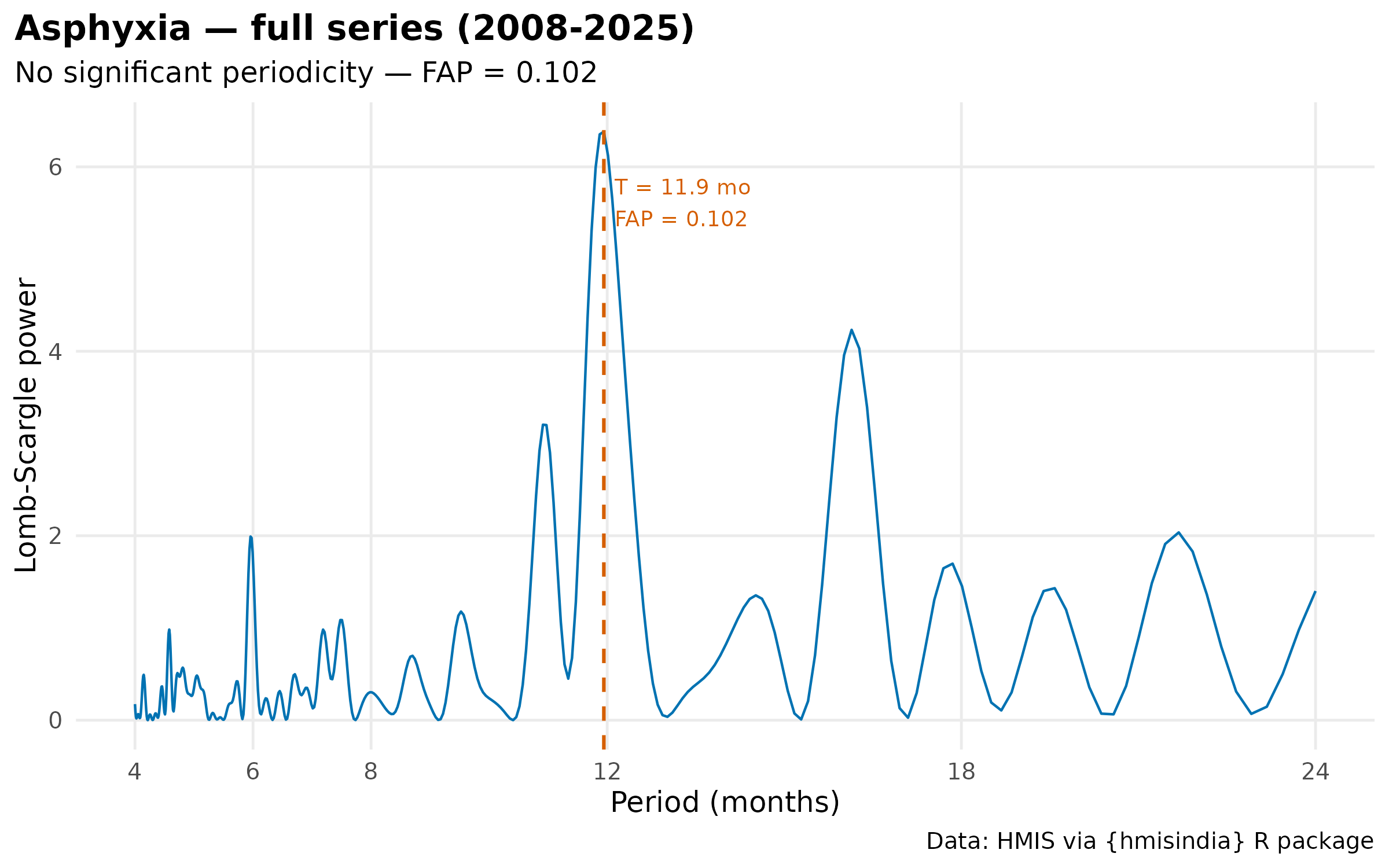
plot(results[["Asphyxia_new"]]$result) +
labs(title = "Asphyxia — new series only (2018-2024)") +
theme_hmis() + labs_hmis()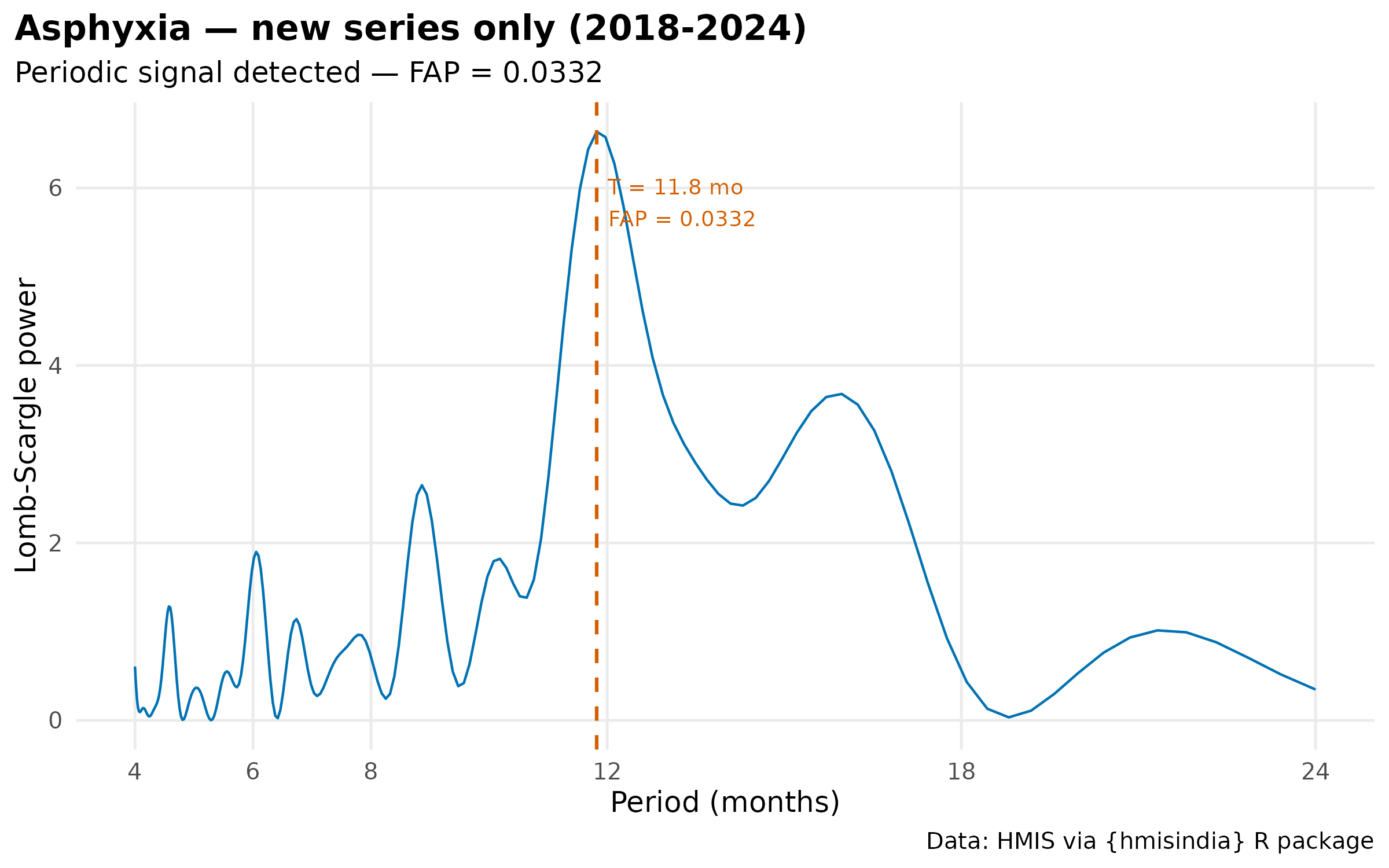
plot(results[["Sepsis_full"]]$result) +
labs(title = "Sepsis — full series (2008-2025)") +
theme_hmis() + labs_hmis()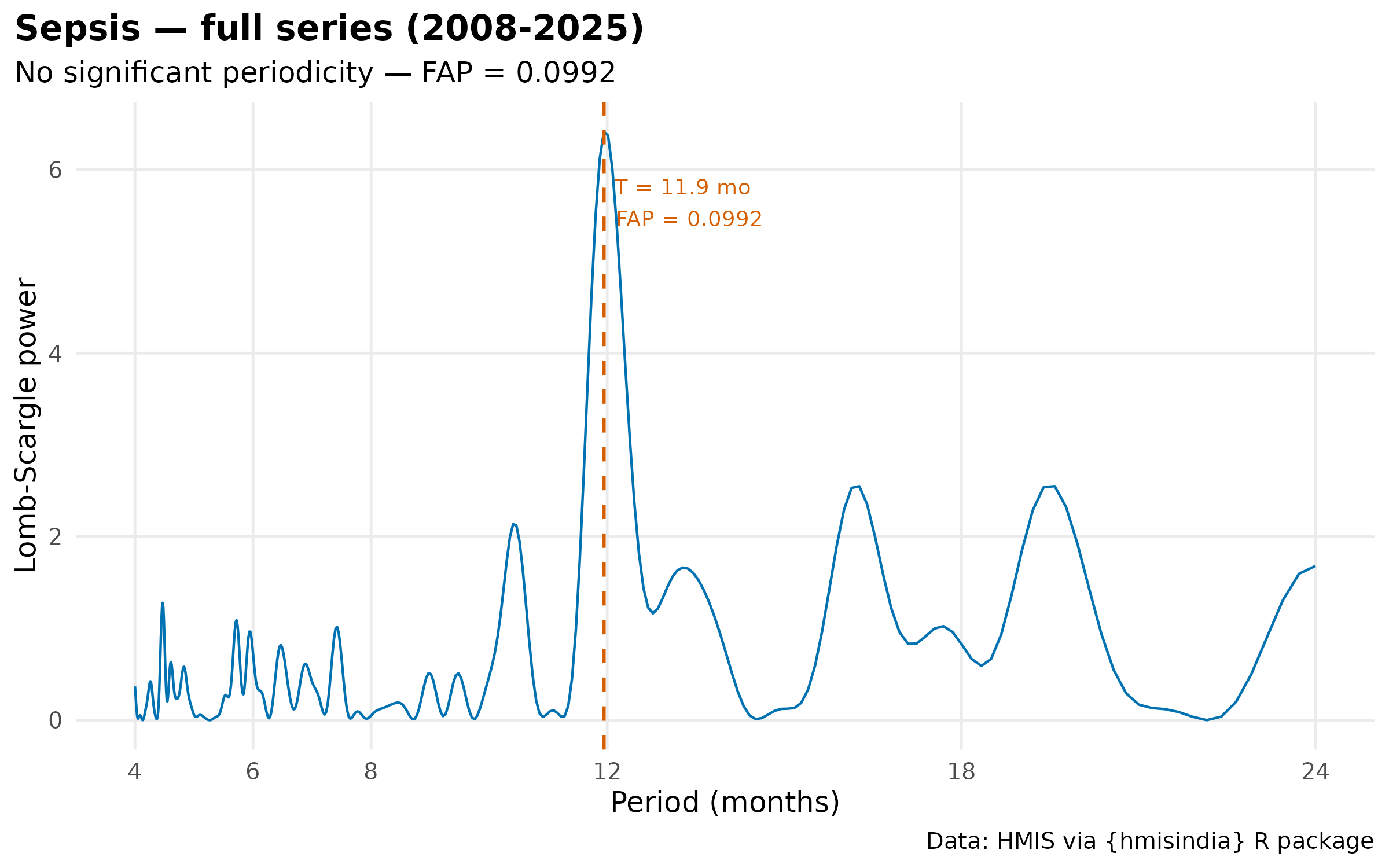
plot(results[["Sepsis_new"]]$result) +
labs(title = "Sepsis — new series only (2018-2024)") +
theme_hmis() + labs_hmis()